Image Contest 2025- BINA 2025 Light Microscopy Image Contest
We are so thankful for all our 2025 Image Contest Contributors!!!
The Top three images are:
- Hexagonal Vision: Compound Eye and Sensory Crown – Frederic Bonnet
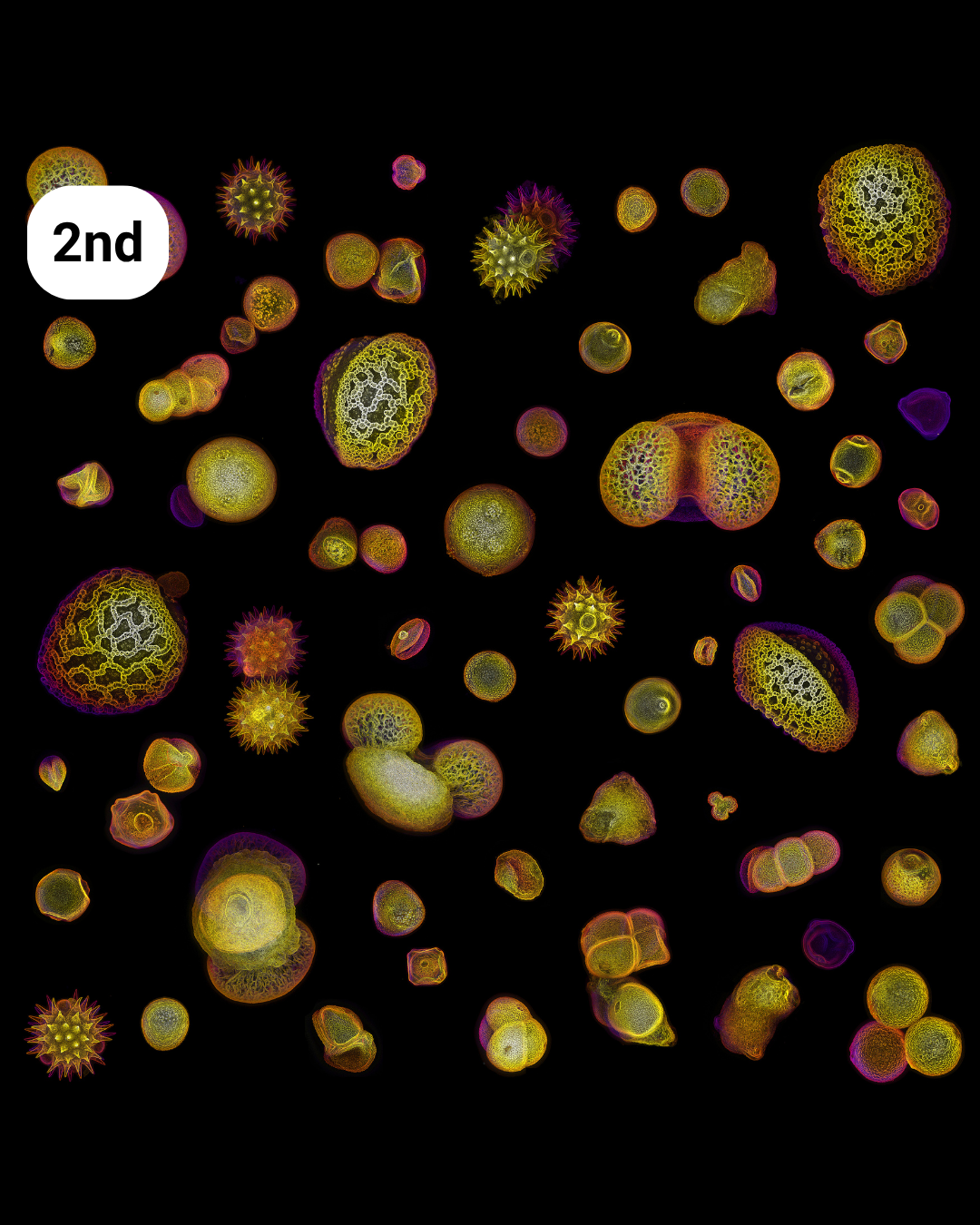
- Allergy Season – Nat Prunet
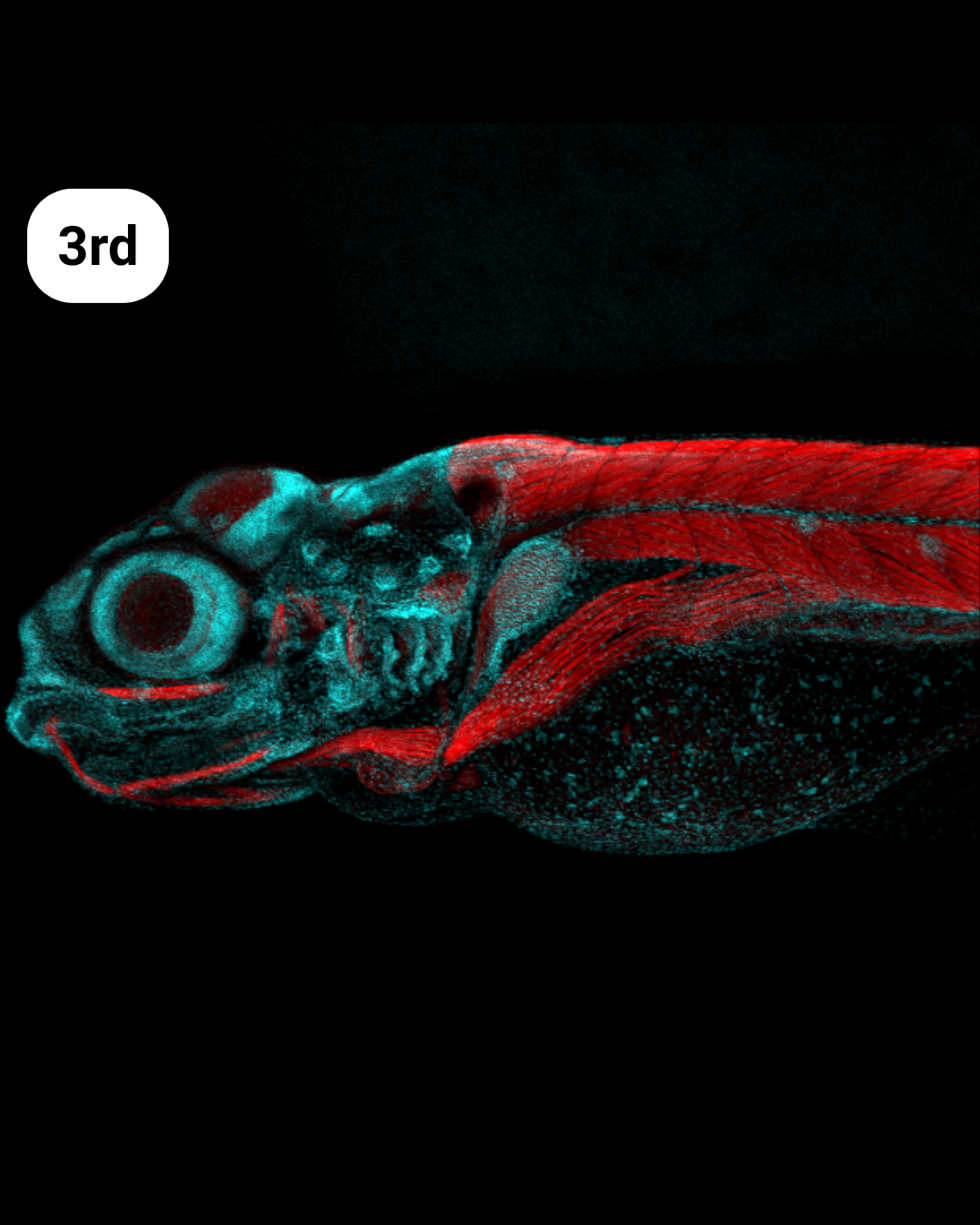
- Blue, Red and Fish – Arlen Ramírez Corona
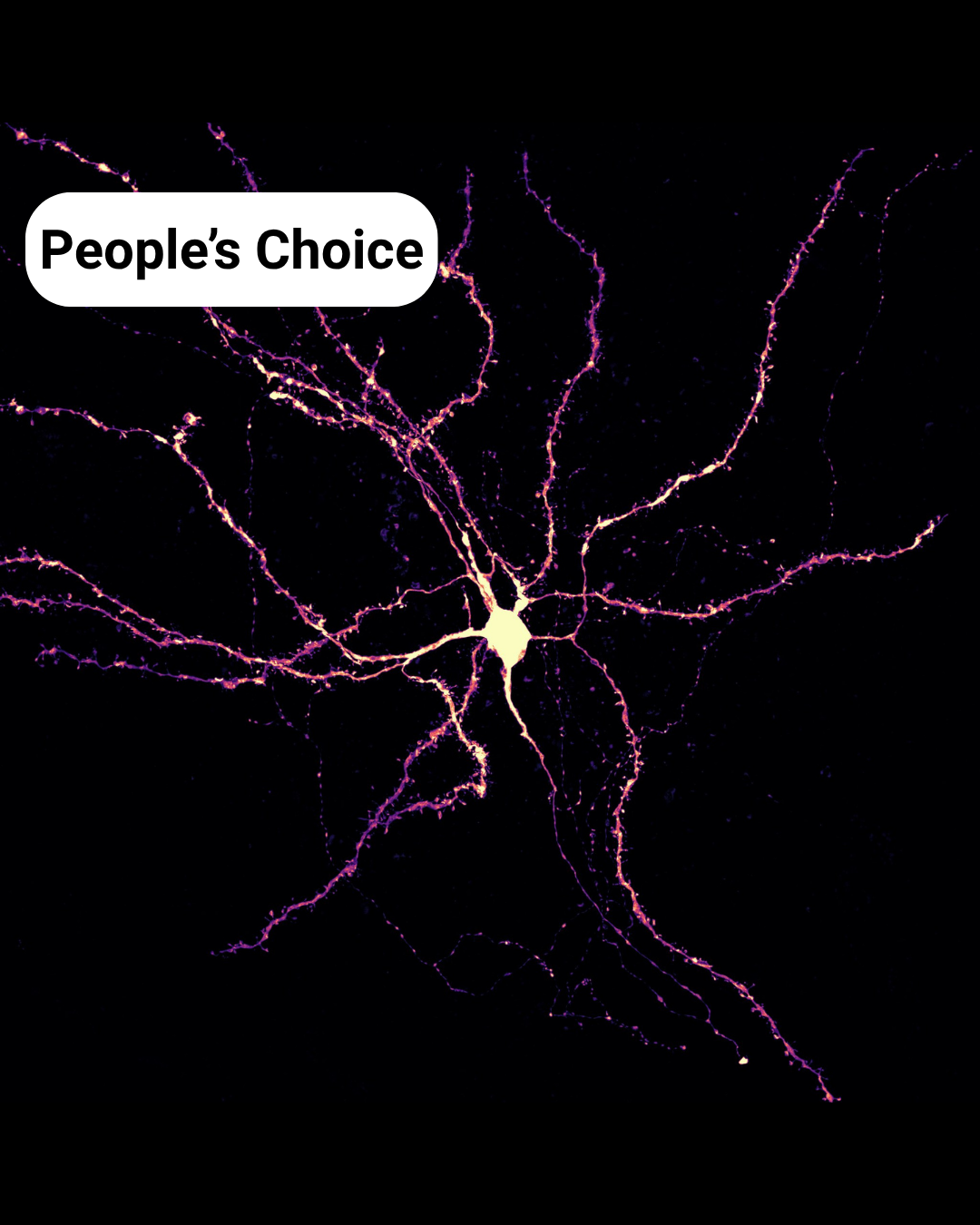
The People’s Choice Award is: Dentritic constellation – Mahgol Darvishmolla
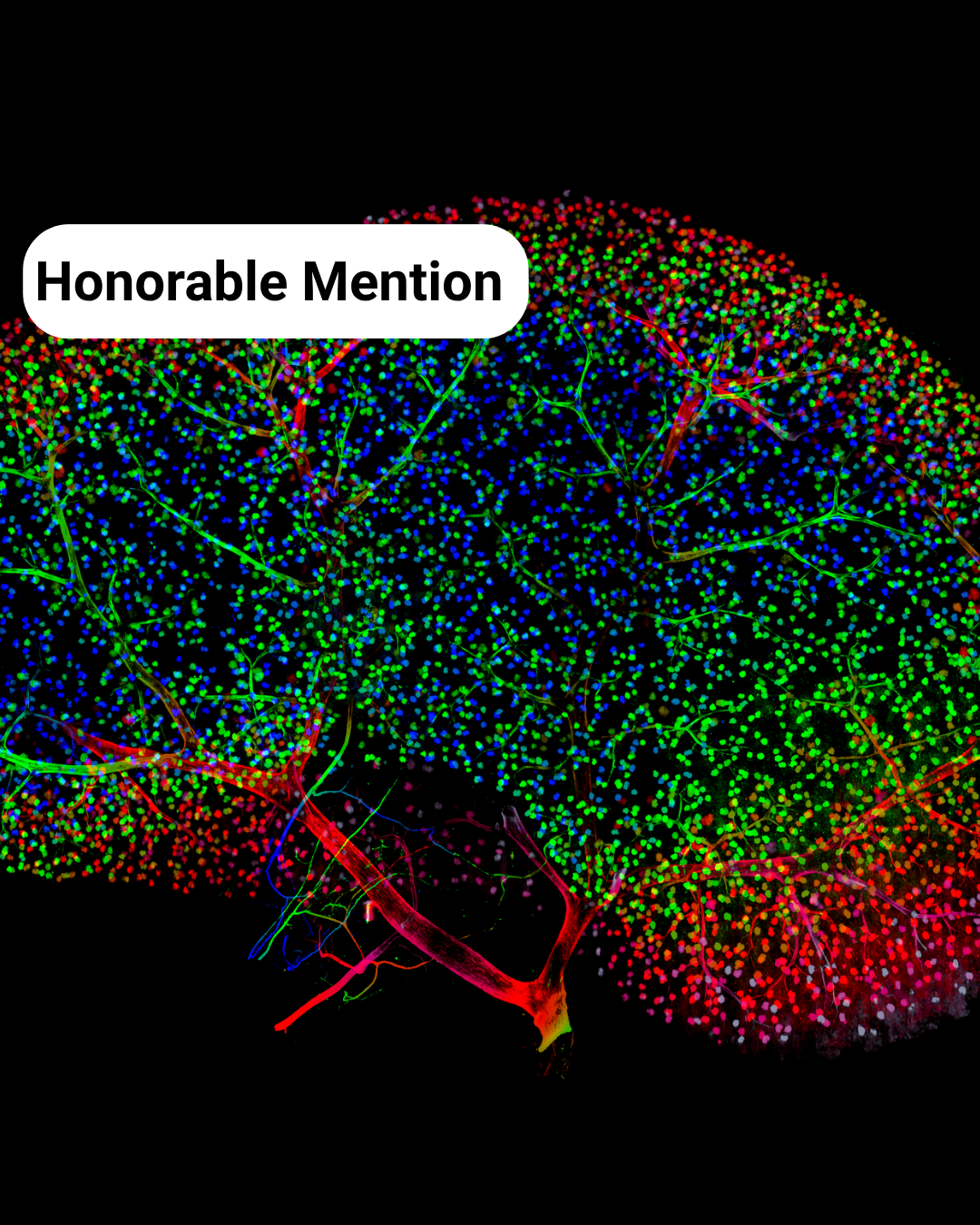
Honorable Mentions (alpha order)
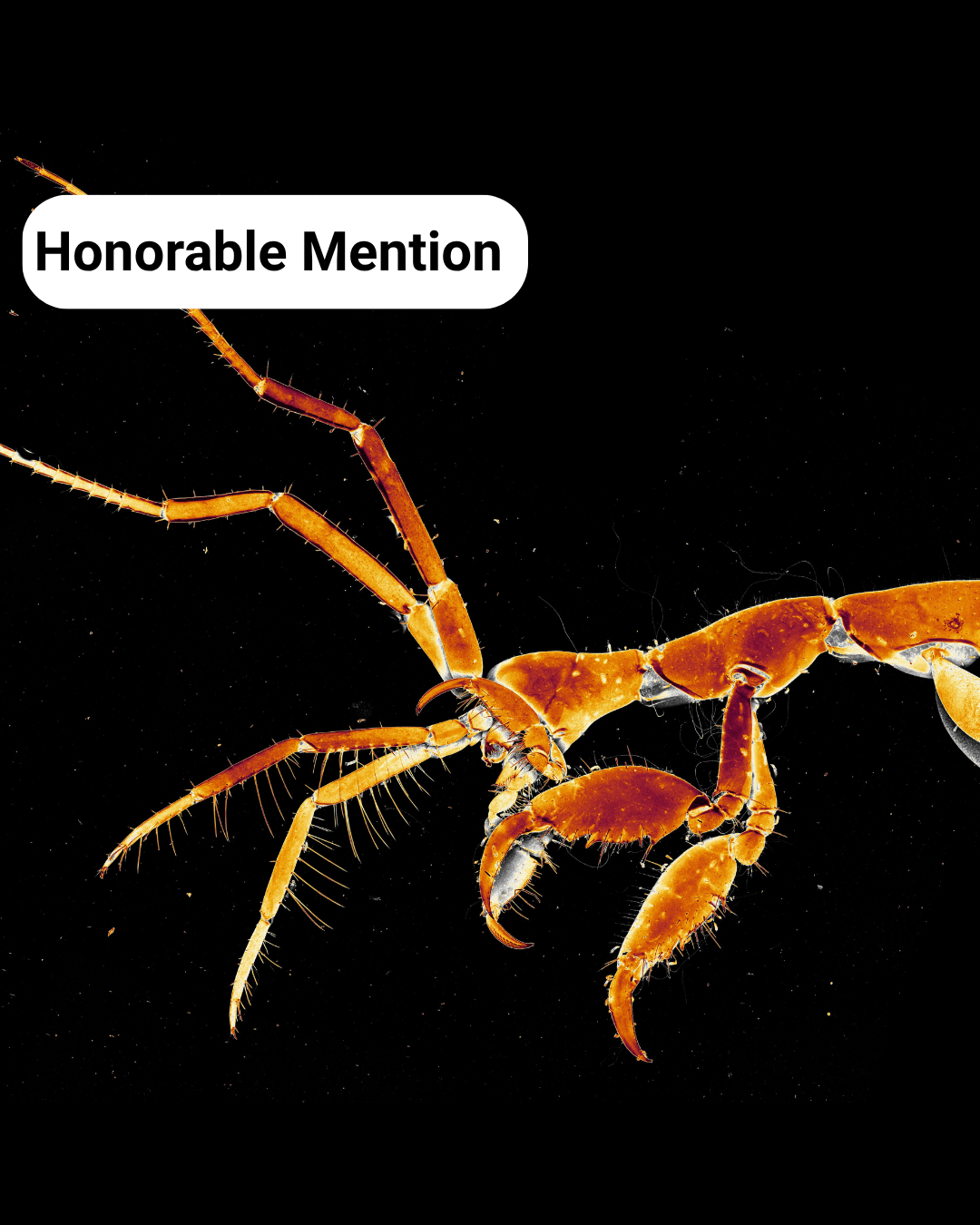
- Textures of a Skeleton Shrimp – Caroline Hoppe
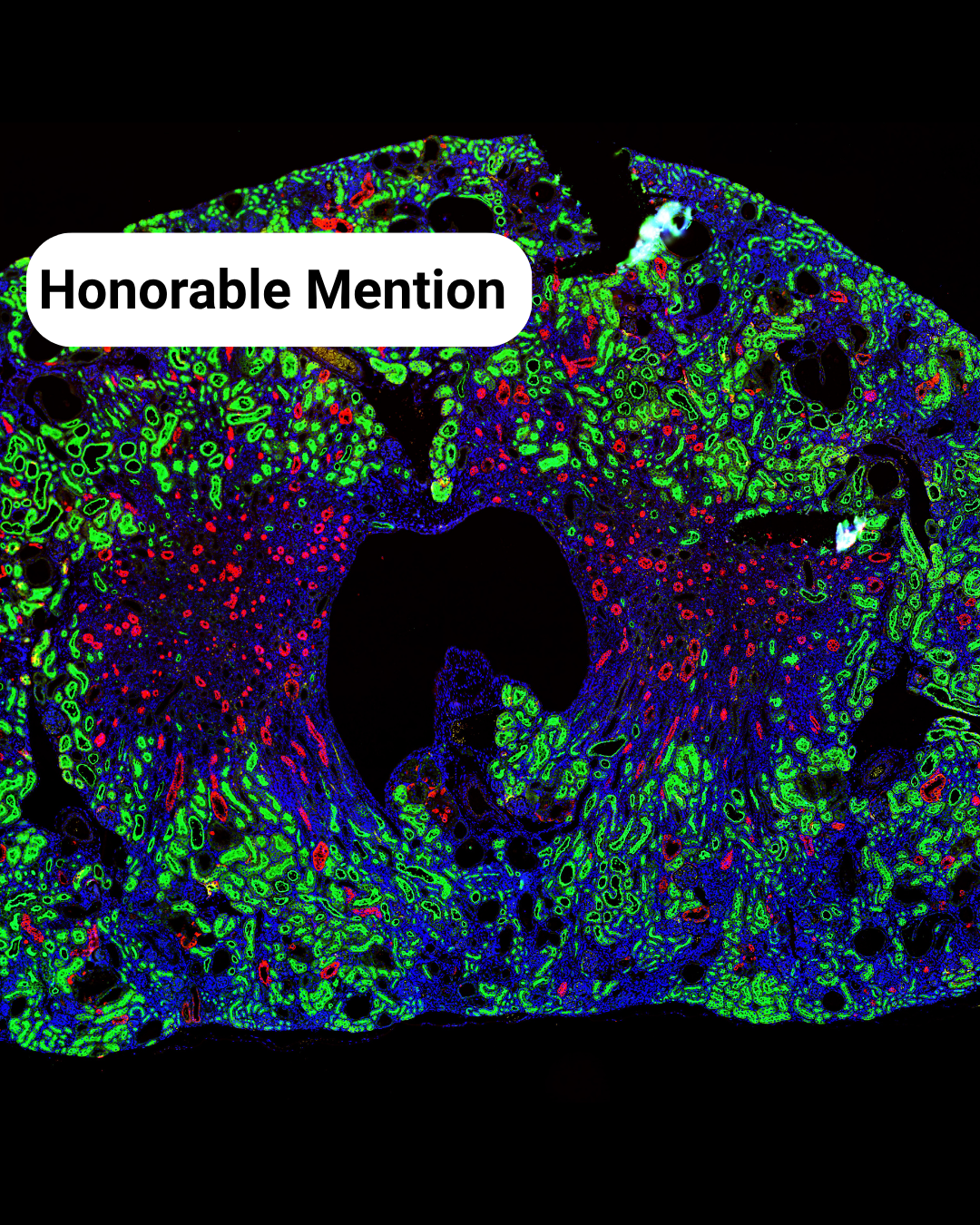
- Mouse Kidney Disco – Chetan Poudel
- The Kidney Landscape** – Meriam Shabbar
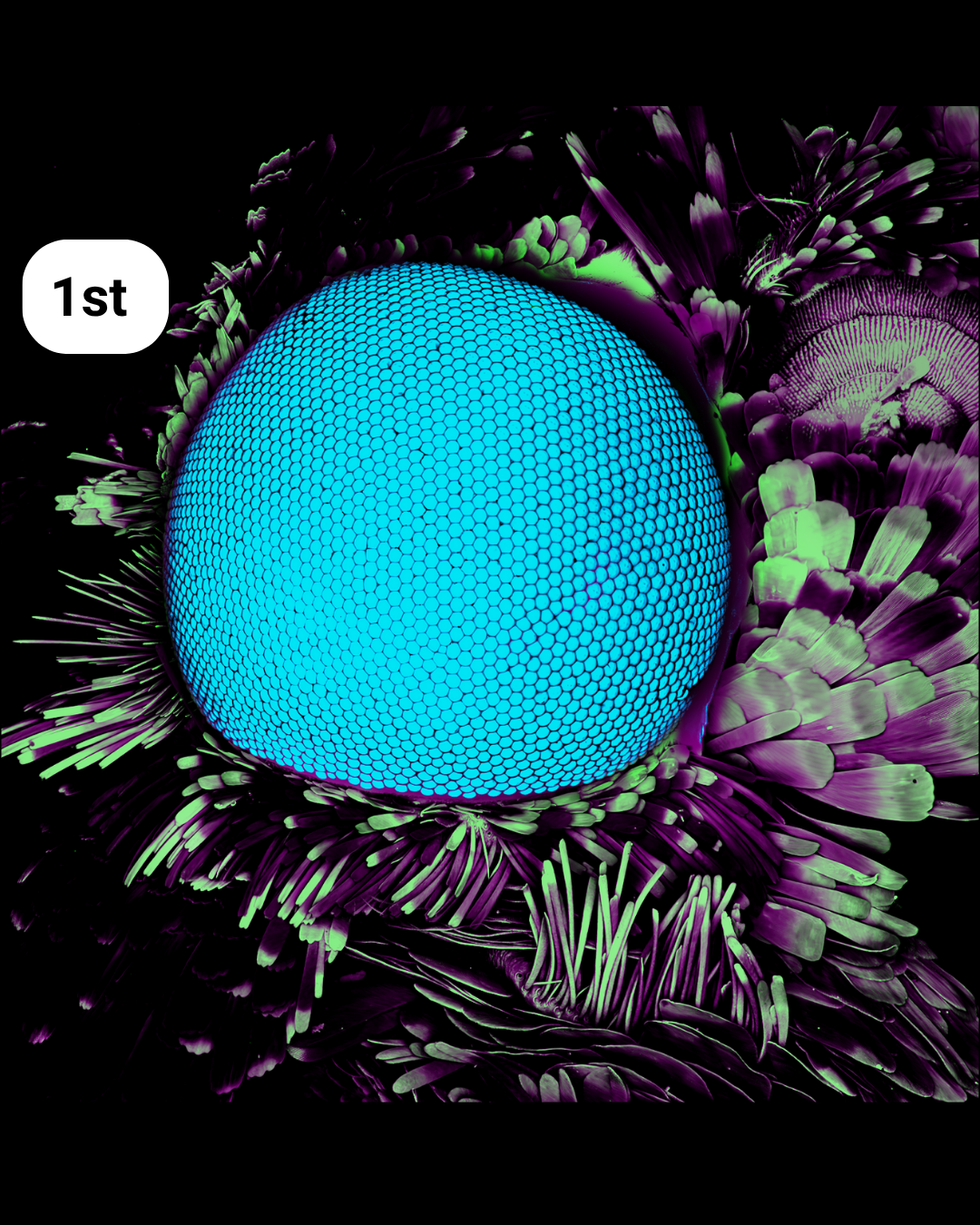

Hexagonal Vision: Compound Eye and Sensory Crown– Frederic Bonnet
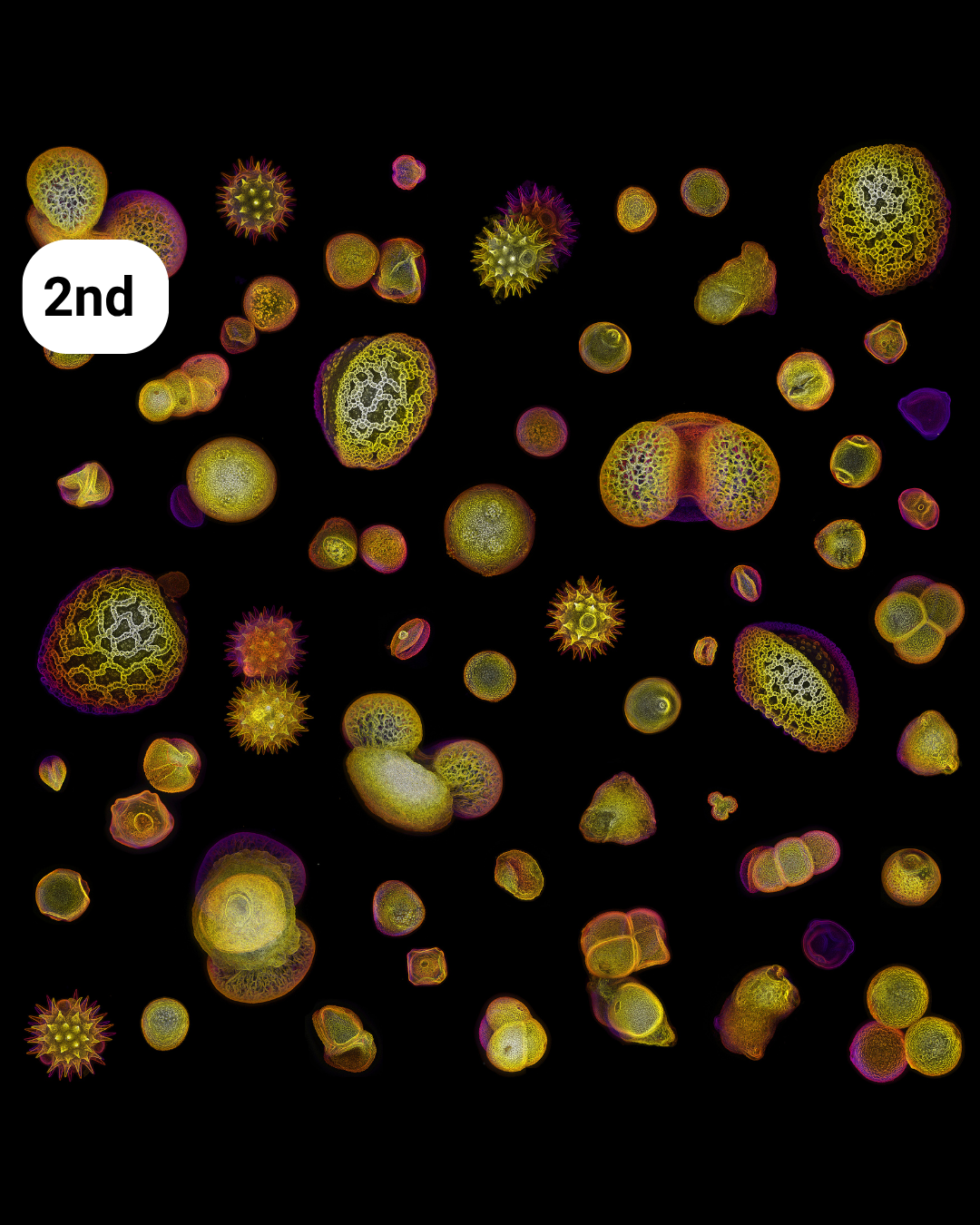
Allergy season– Nat Prunet
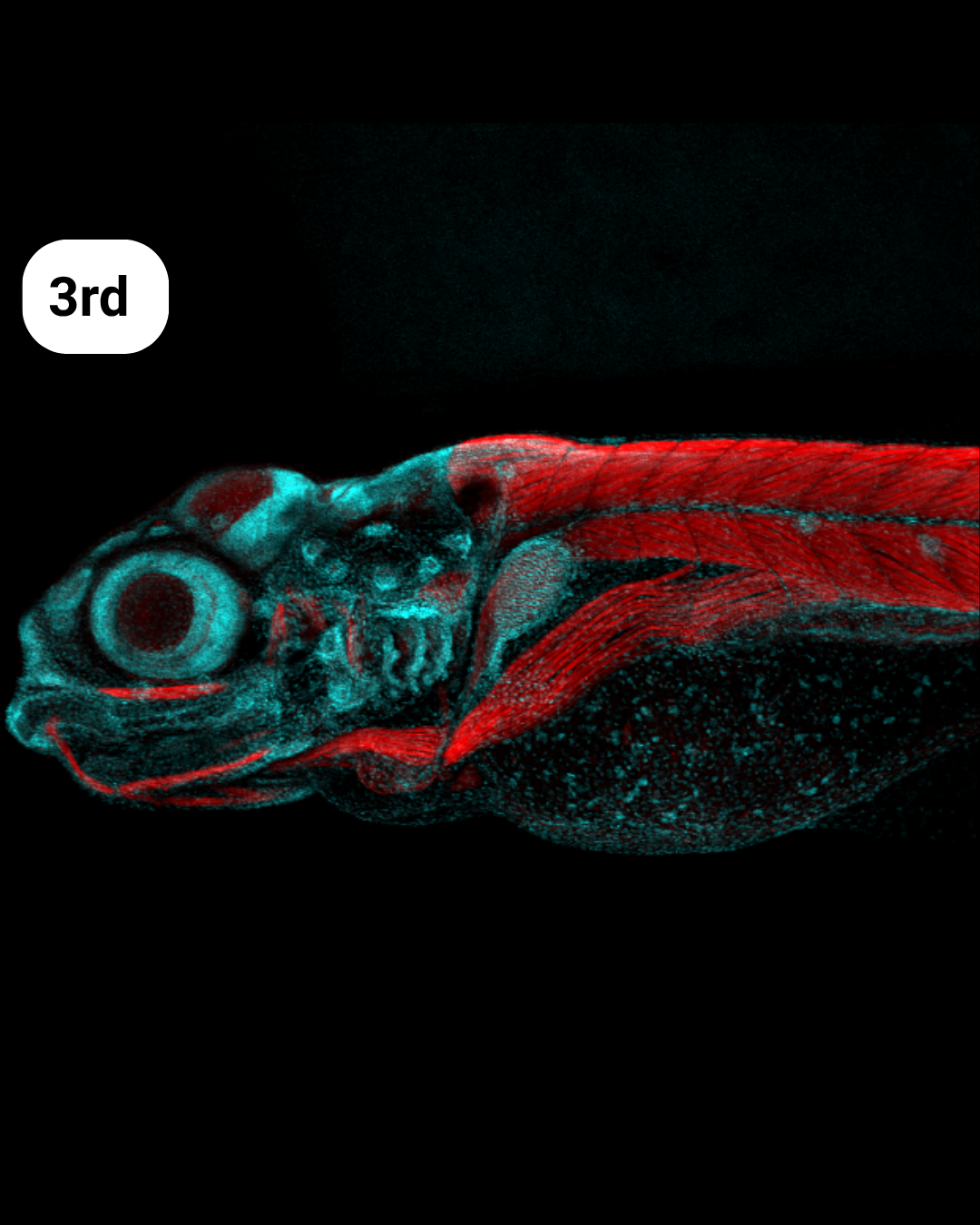
Blue, Red and Fish- Arlen Ramírez Corona
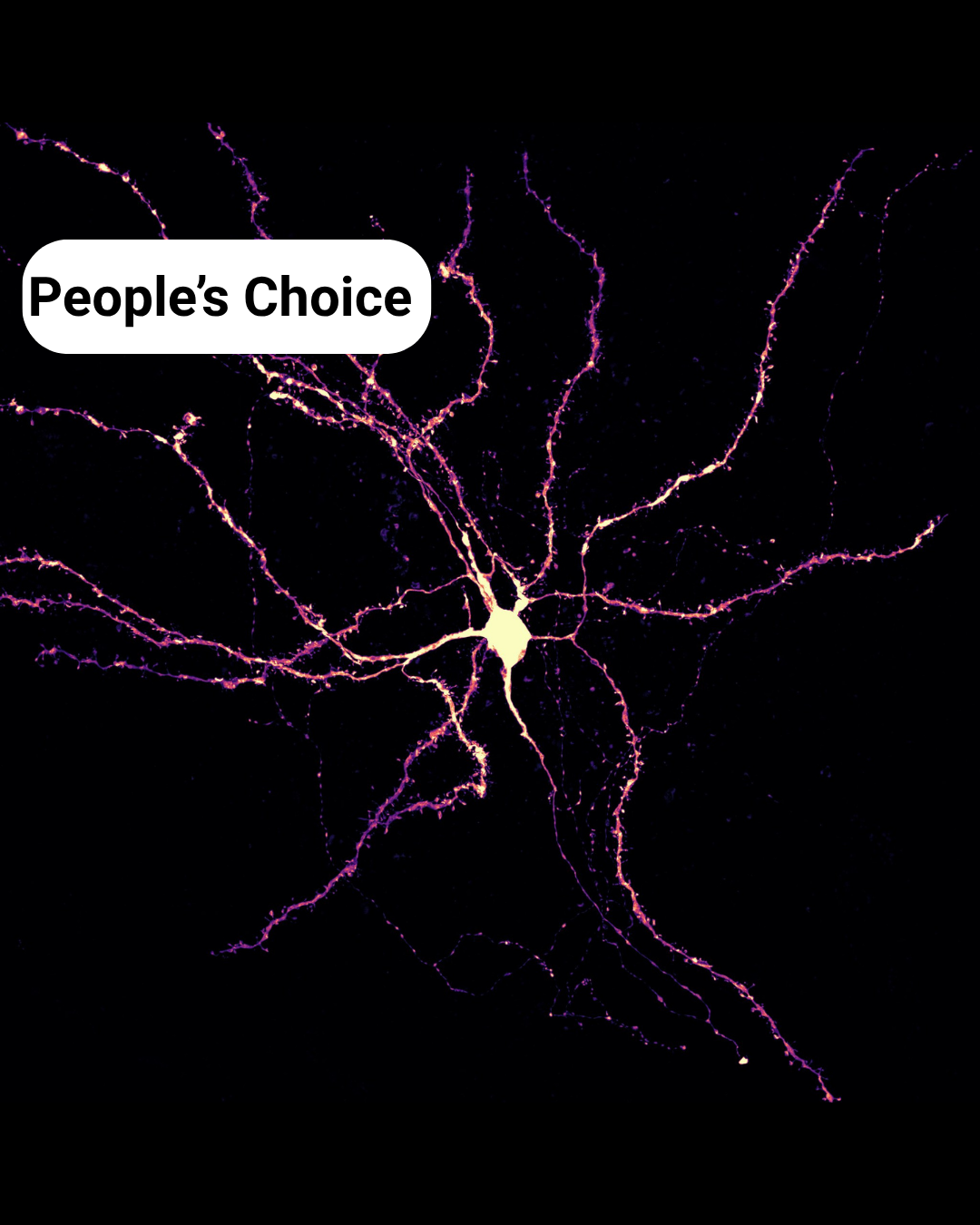
Dentritic constellation– Mahgol Darvishmolla
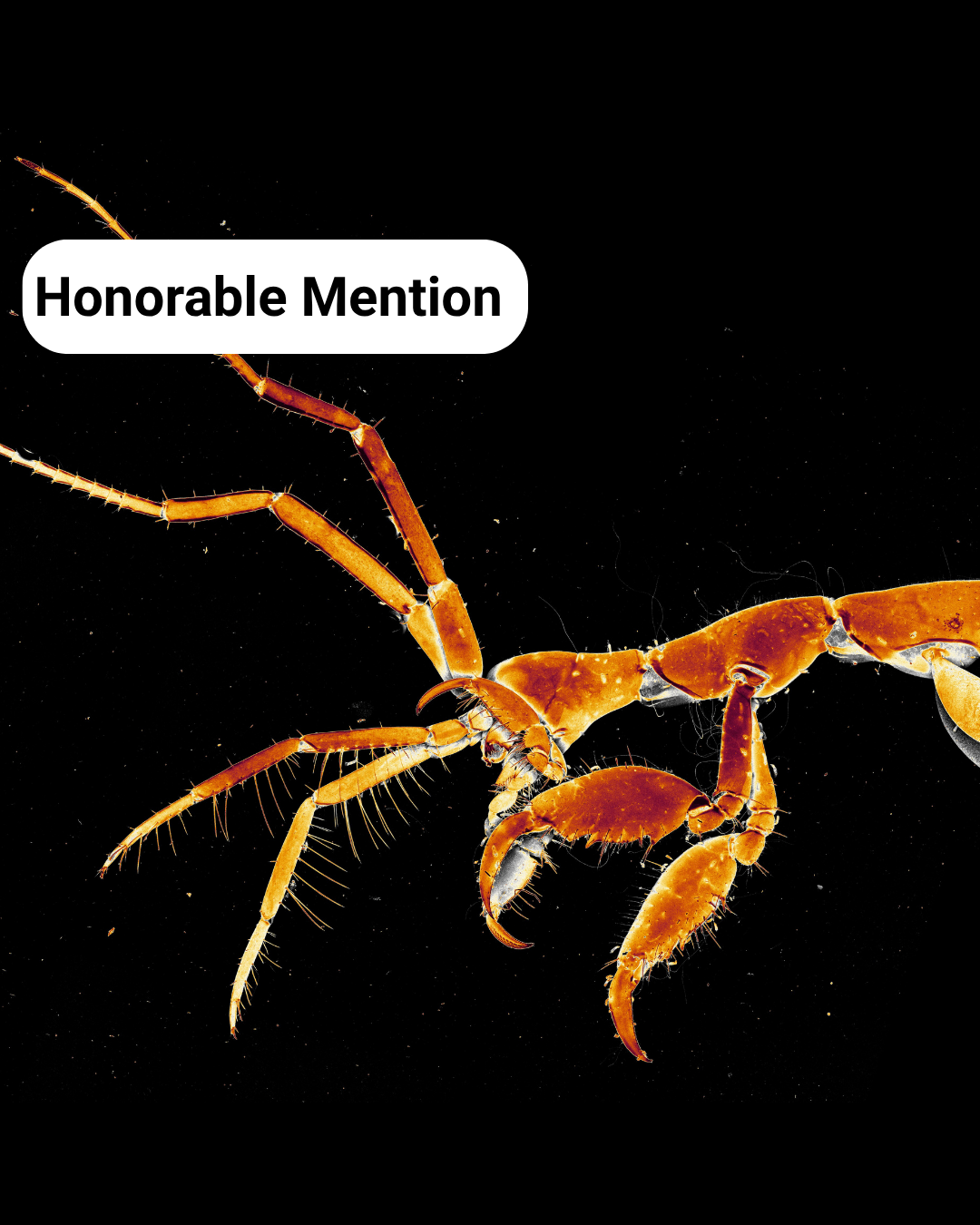

Textures of a Skeleton Shrimp– Caroline Hoppe

Mouse Kidney Disco- Chetan Poudel
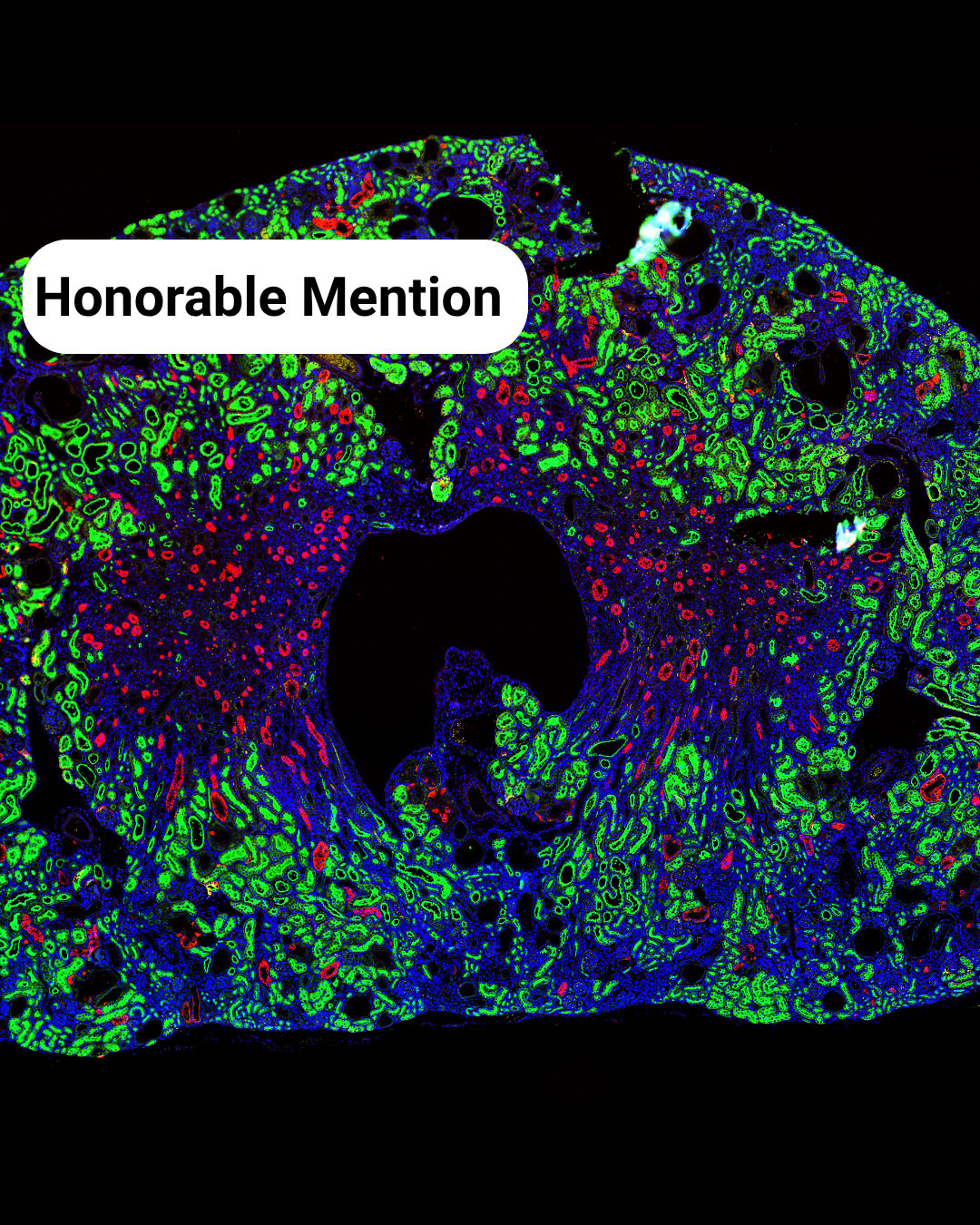
The Kidney Landscape– Meriam Shabbar
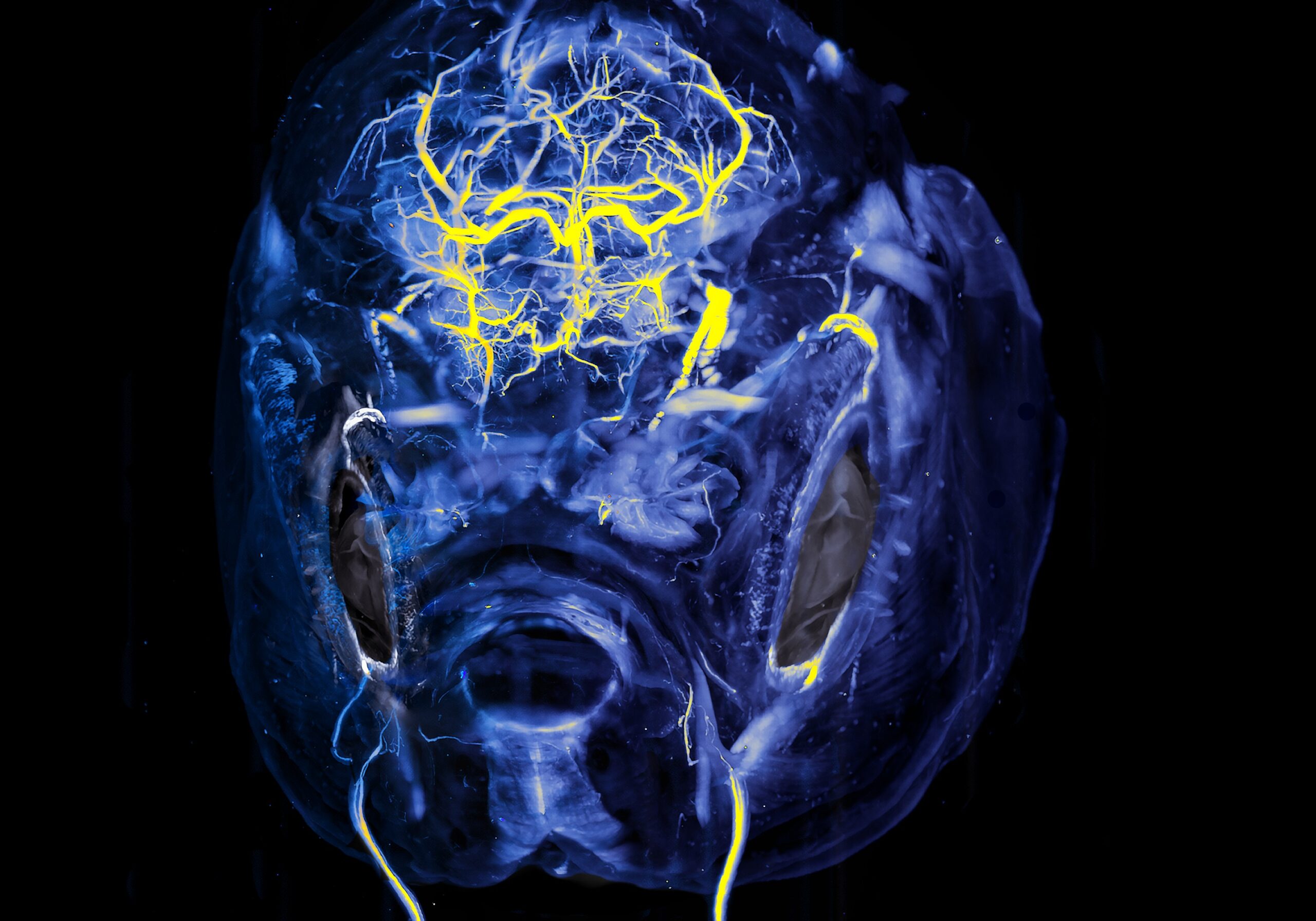
Fish on the Brain- Hannah Somers
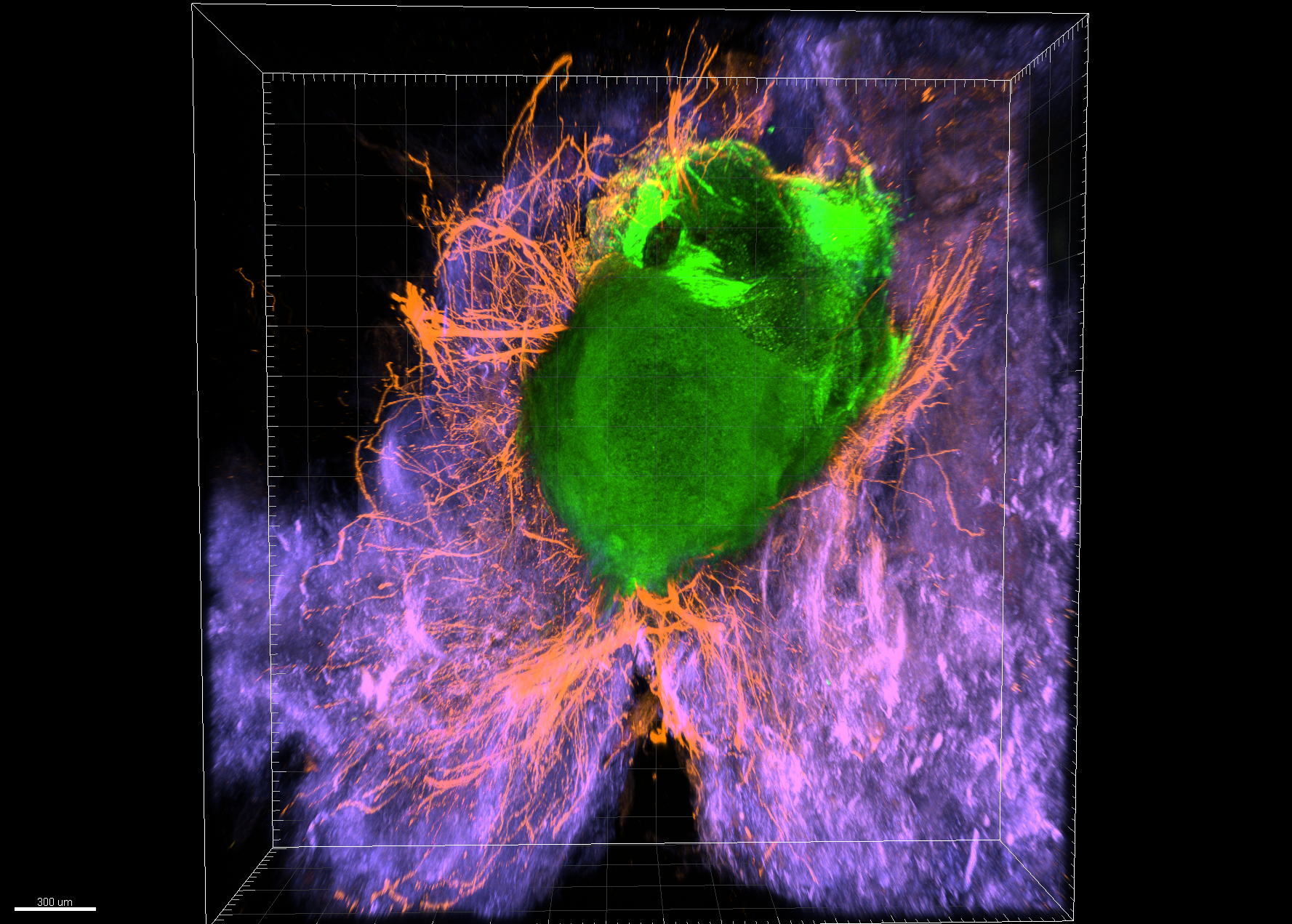
Cortical Organoid in a Bioactive Scaffold– Oscar Carballo-Molina
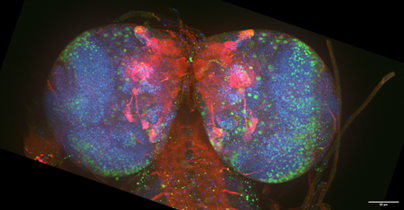
Brain time– Ian Davis
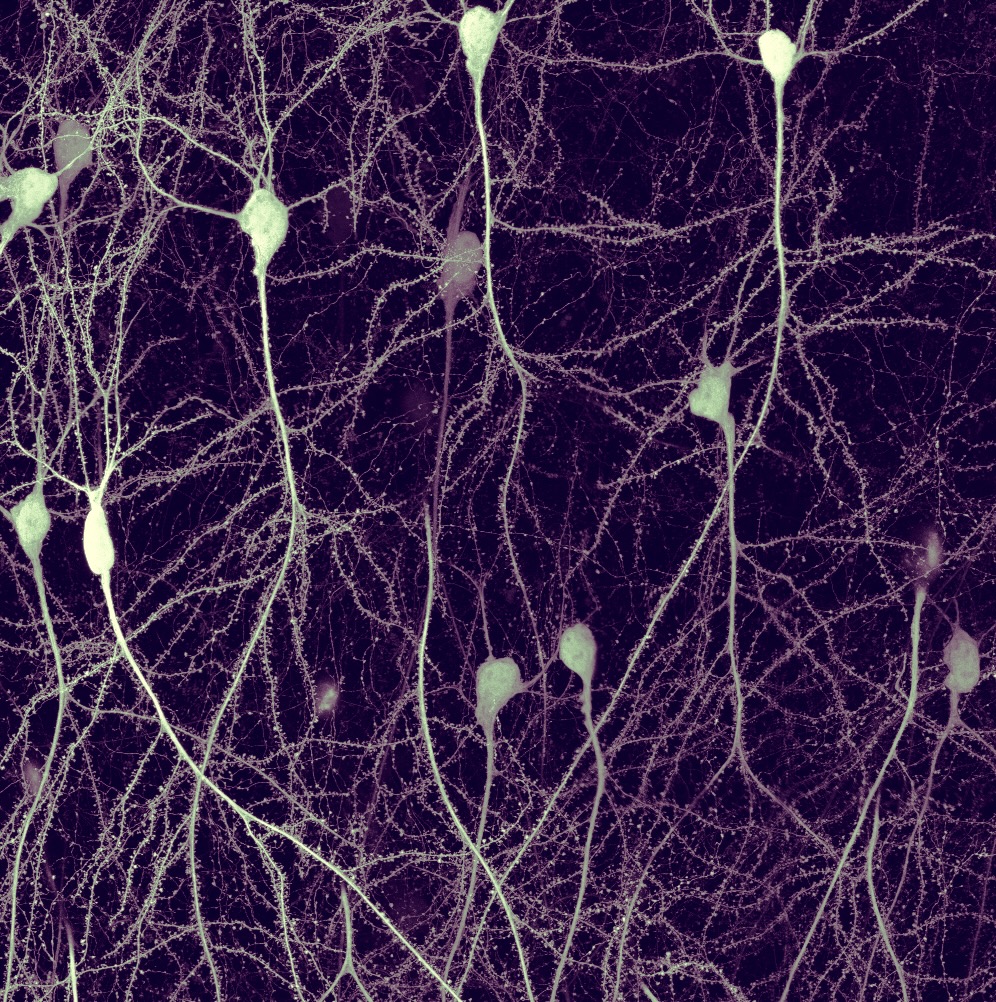
Forest of Memory- Philipp Velicky
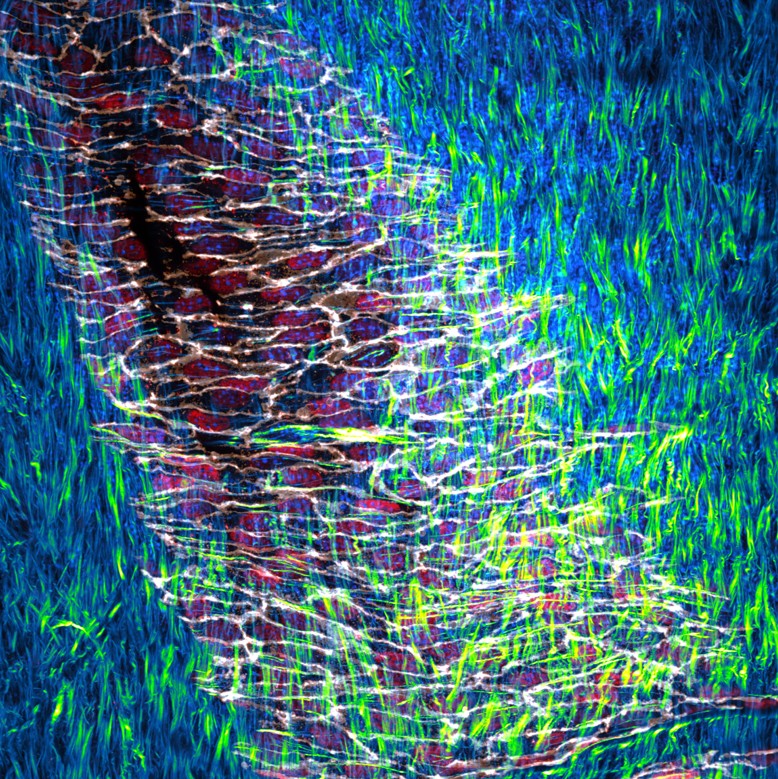
Cobblestone pathway – Team: Mariana De Niz, Jocelyn Salvador, Luisa Iruela-Arispe
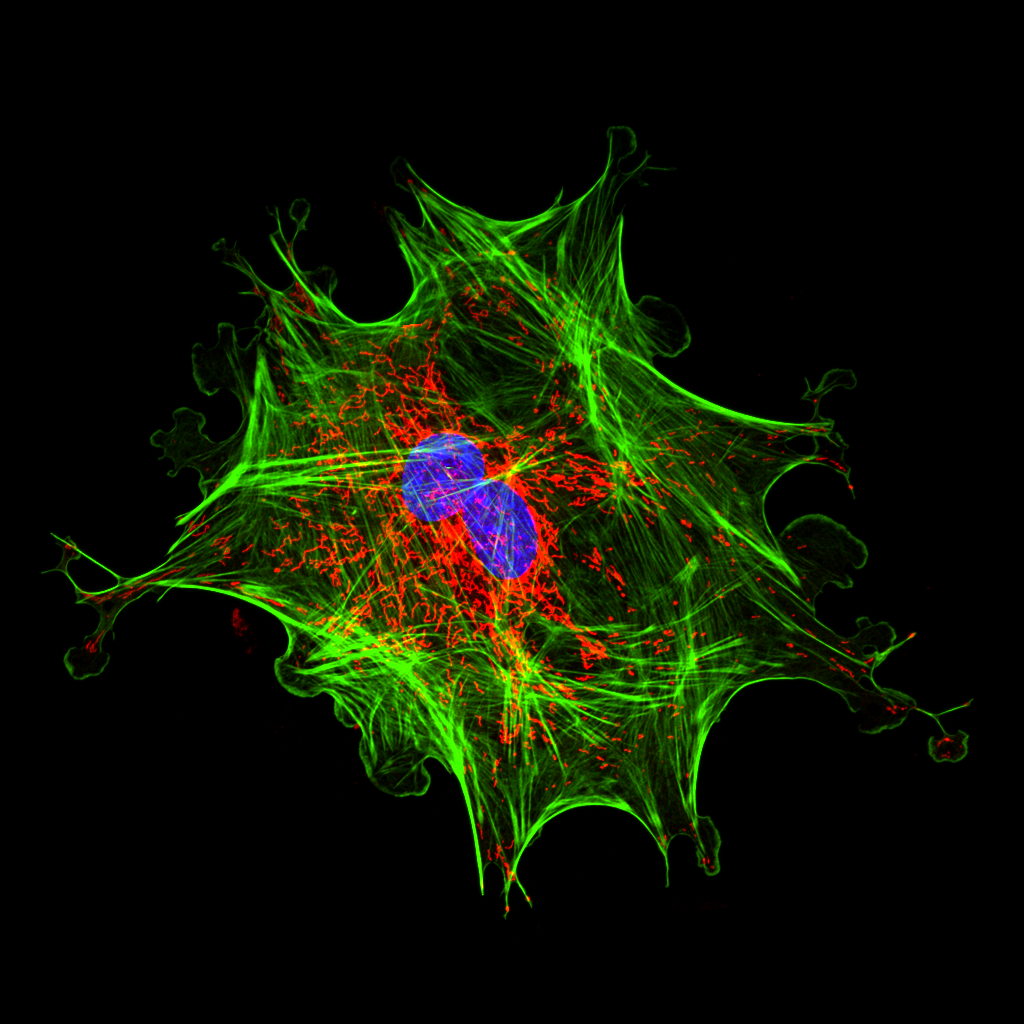
A Starburst of Cellular Life– Sooraj Siini
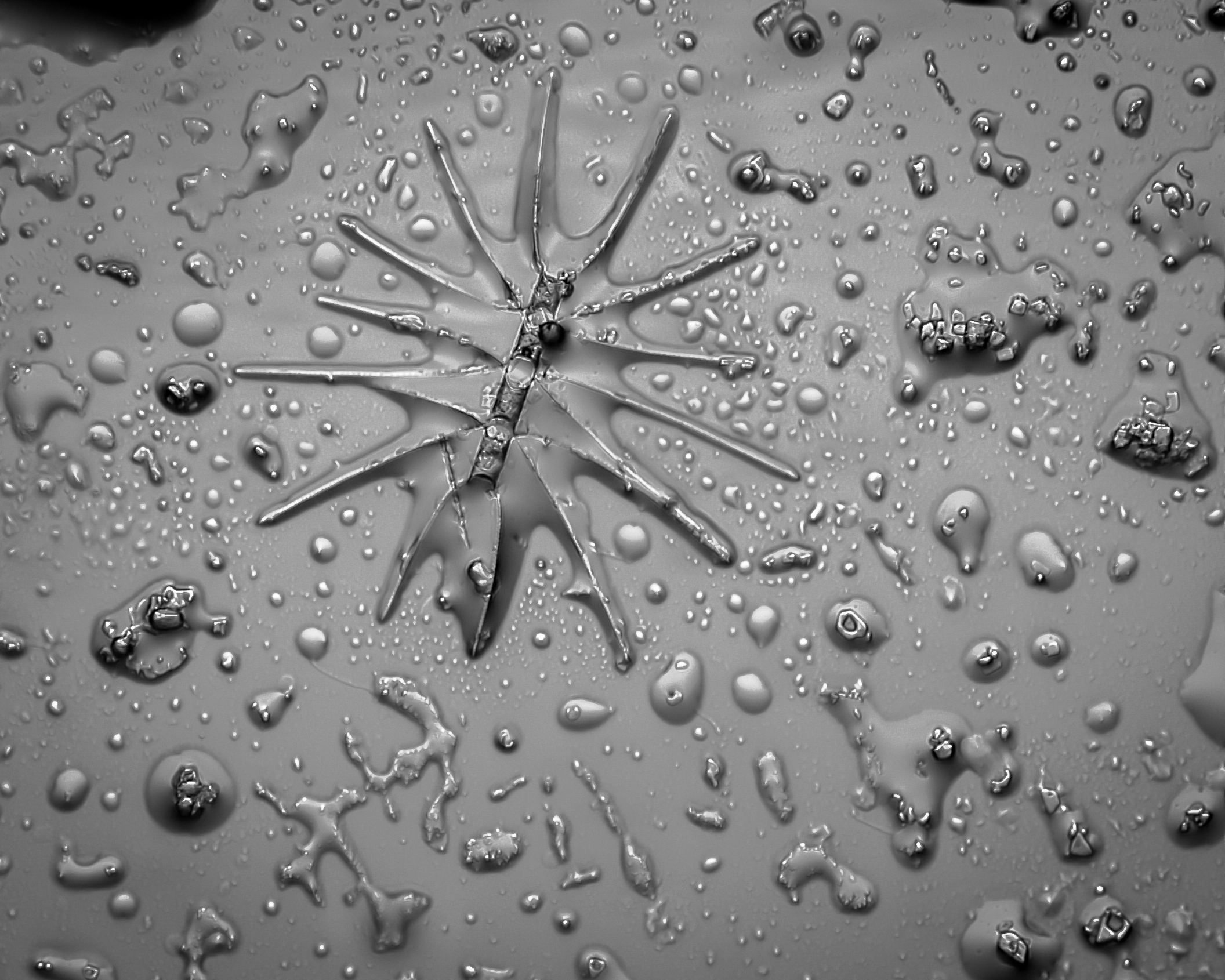

diatom –Jose Manuel Martinez Lopez
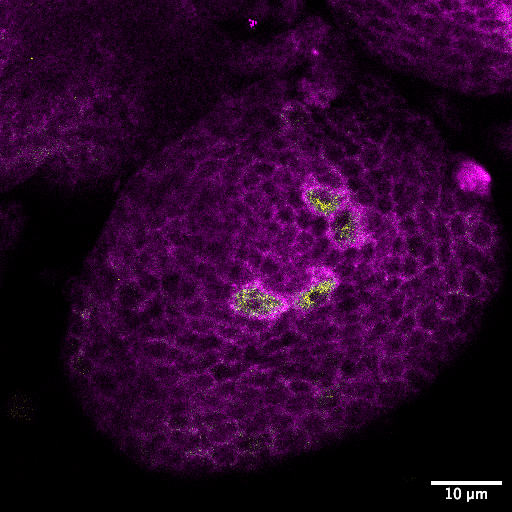
Mushroombody Neuroblasts: Ribosomal Powerhouses of the Fly Brain– Kranti Meher
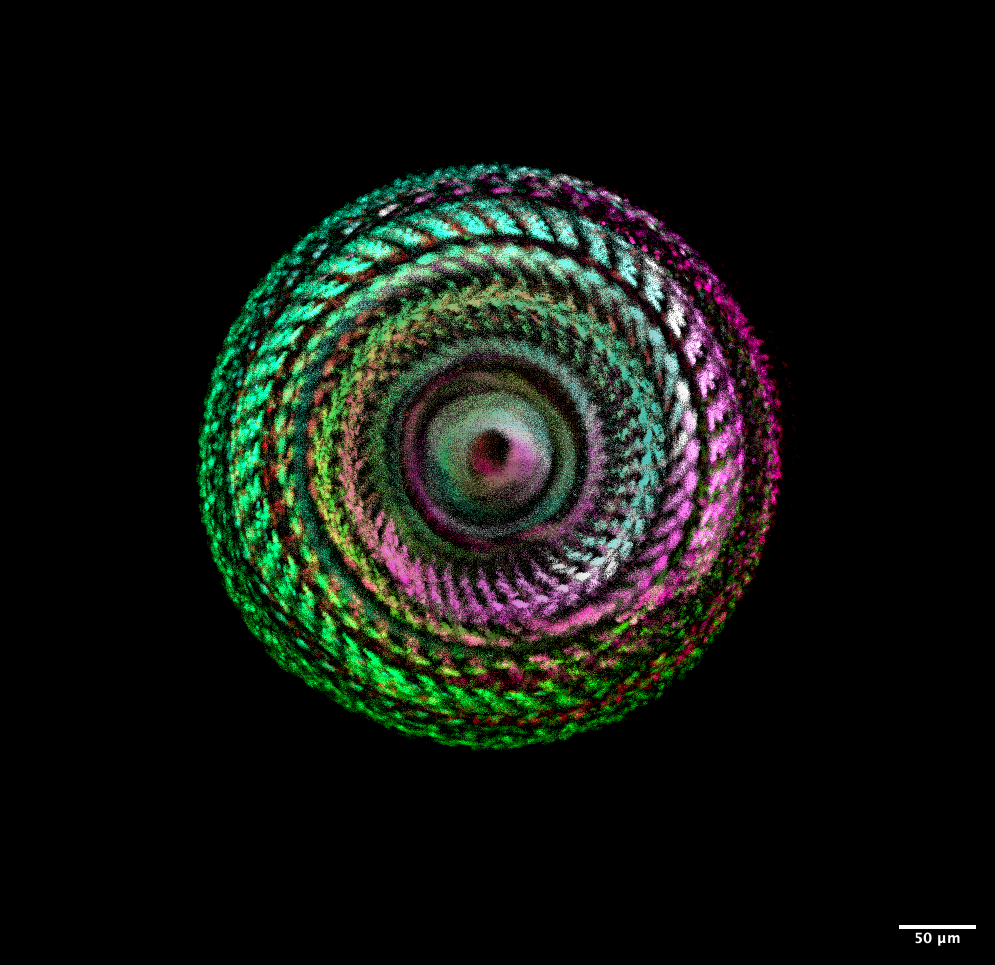

Frozen in Motion– Caroline Hoppe
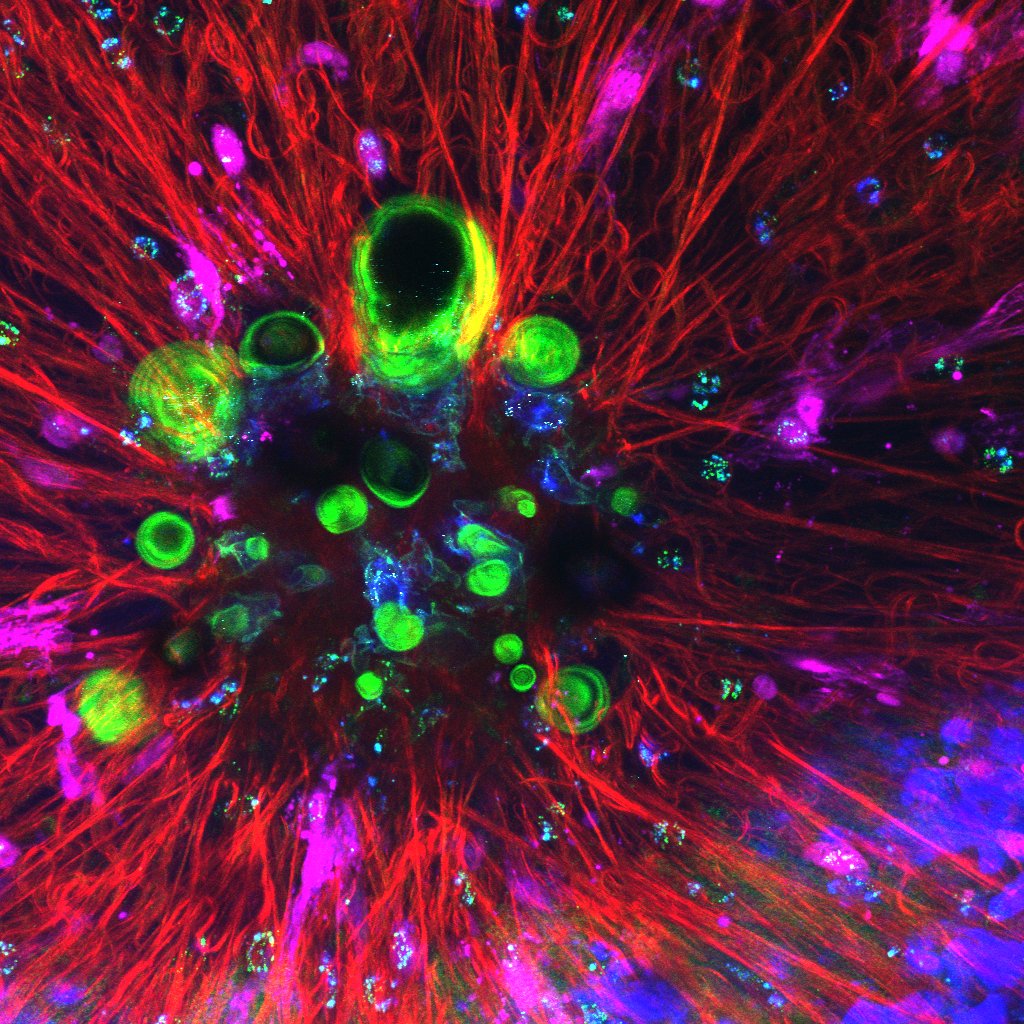
When too much is too much (power)! – Martina Giampetraglia
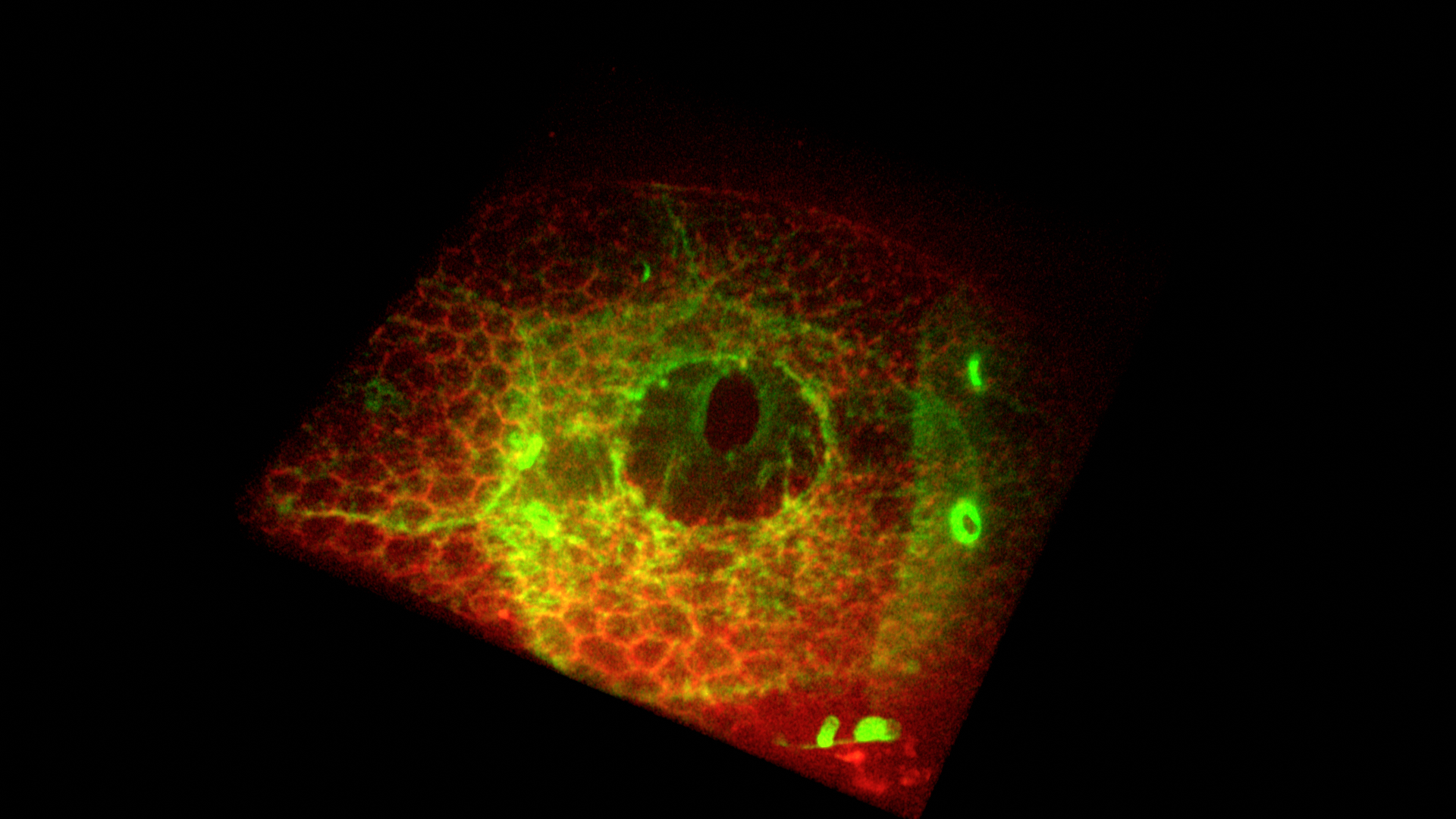
Snapshot of Cells Working to Heal A Wound- Ji Hong Sayo

Image is buffering– Anastasiya Klebanovych
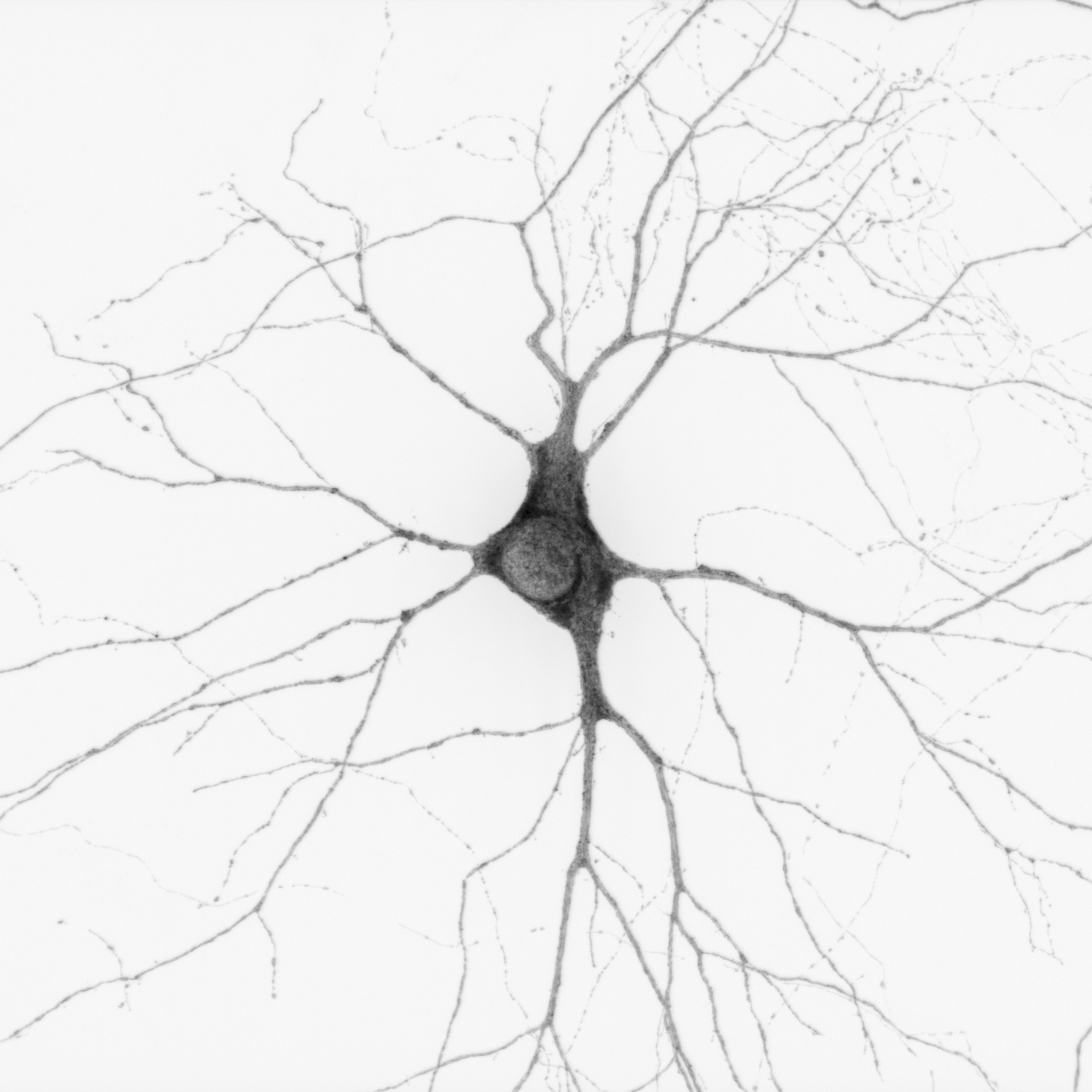
From soma to synapse, the neuronal ER is everywhere!– Cameron Paton
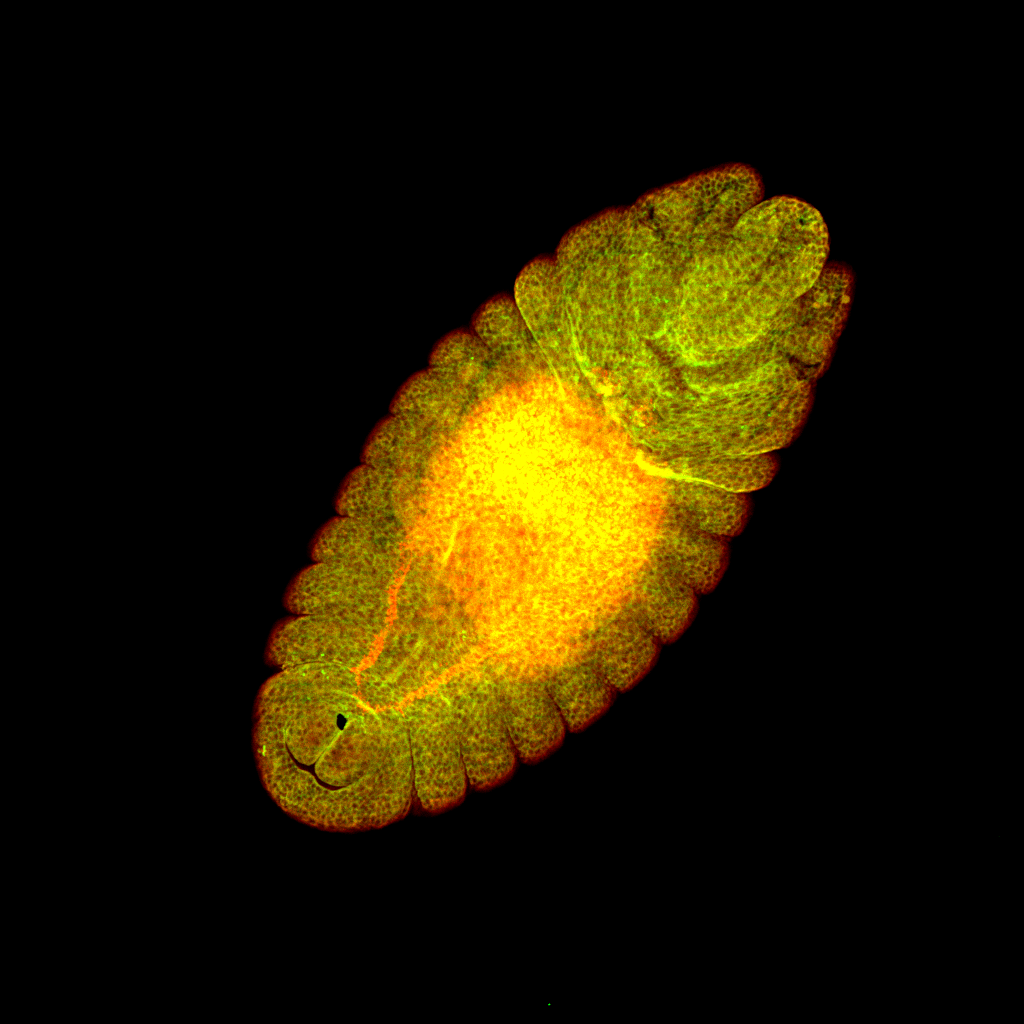
Drosophila melanogaster heart tube development– Rafael Perez Vicente
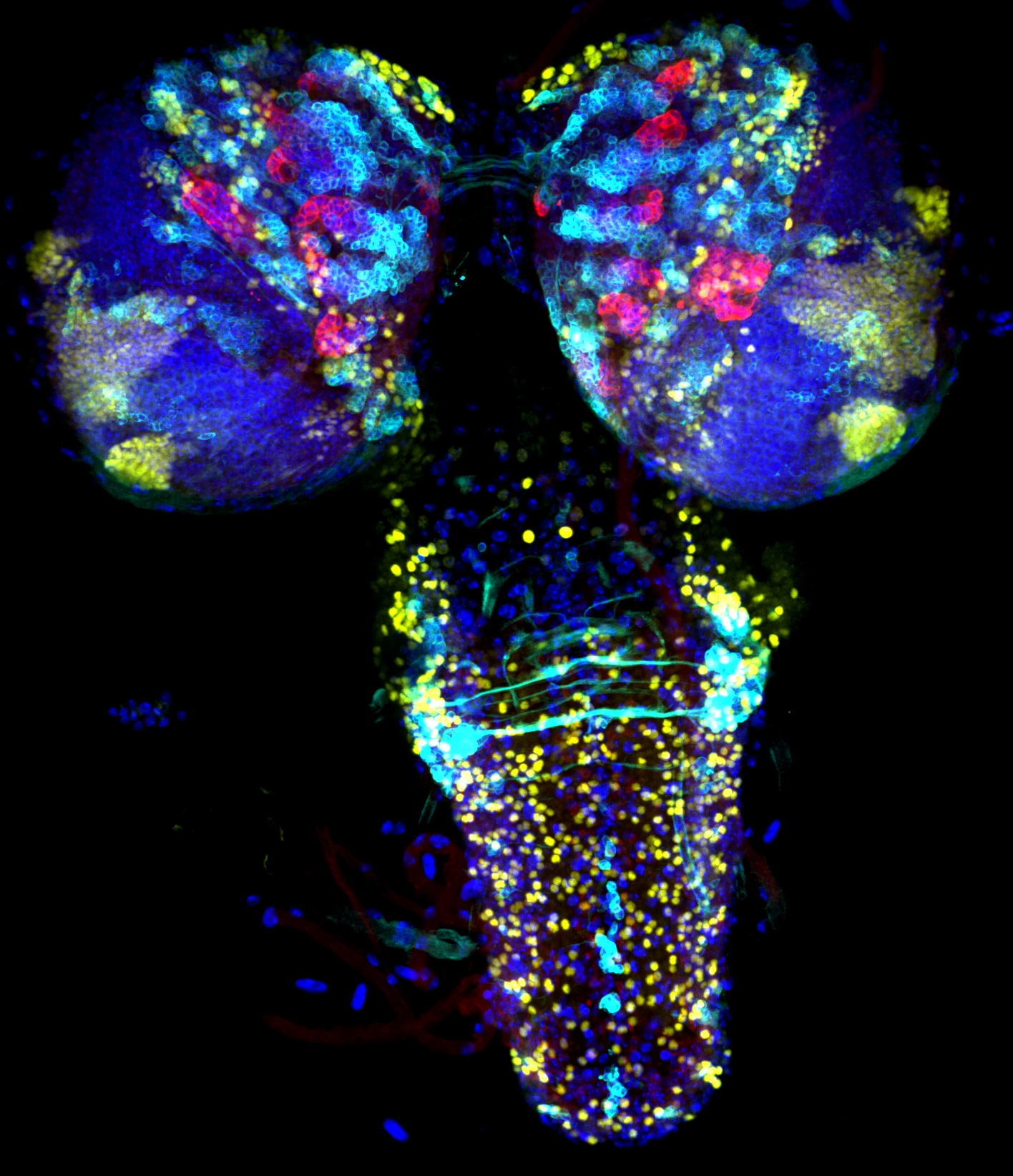
Type two to tango– Travis D. Carney**
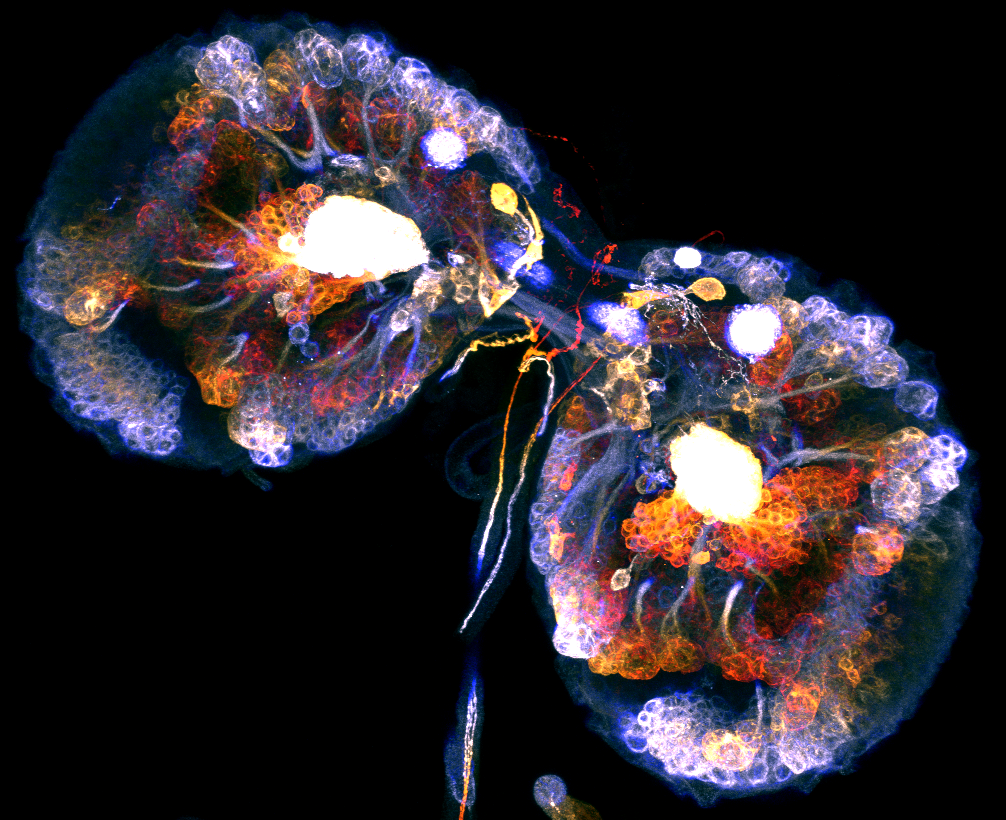
Bal Masque– Travis D. Carney**
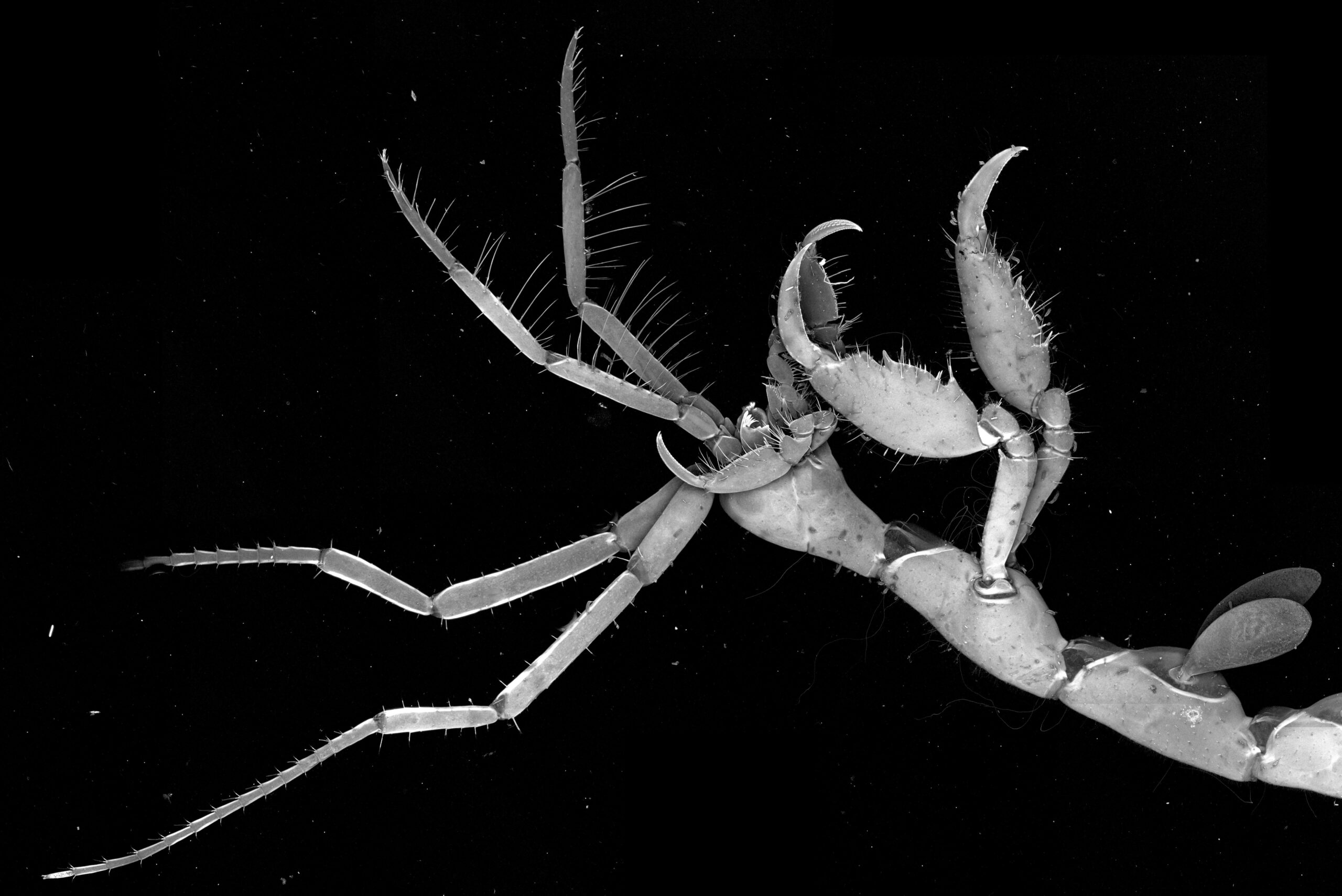

The Grasp of a Skeleton Shrimp– Caroline Hoppe
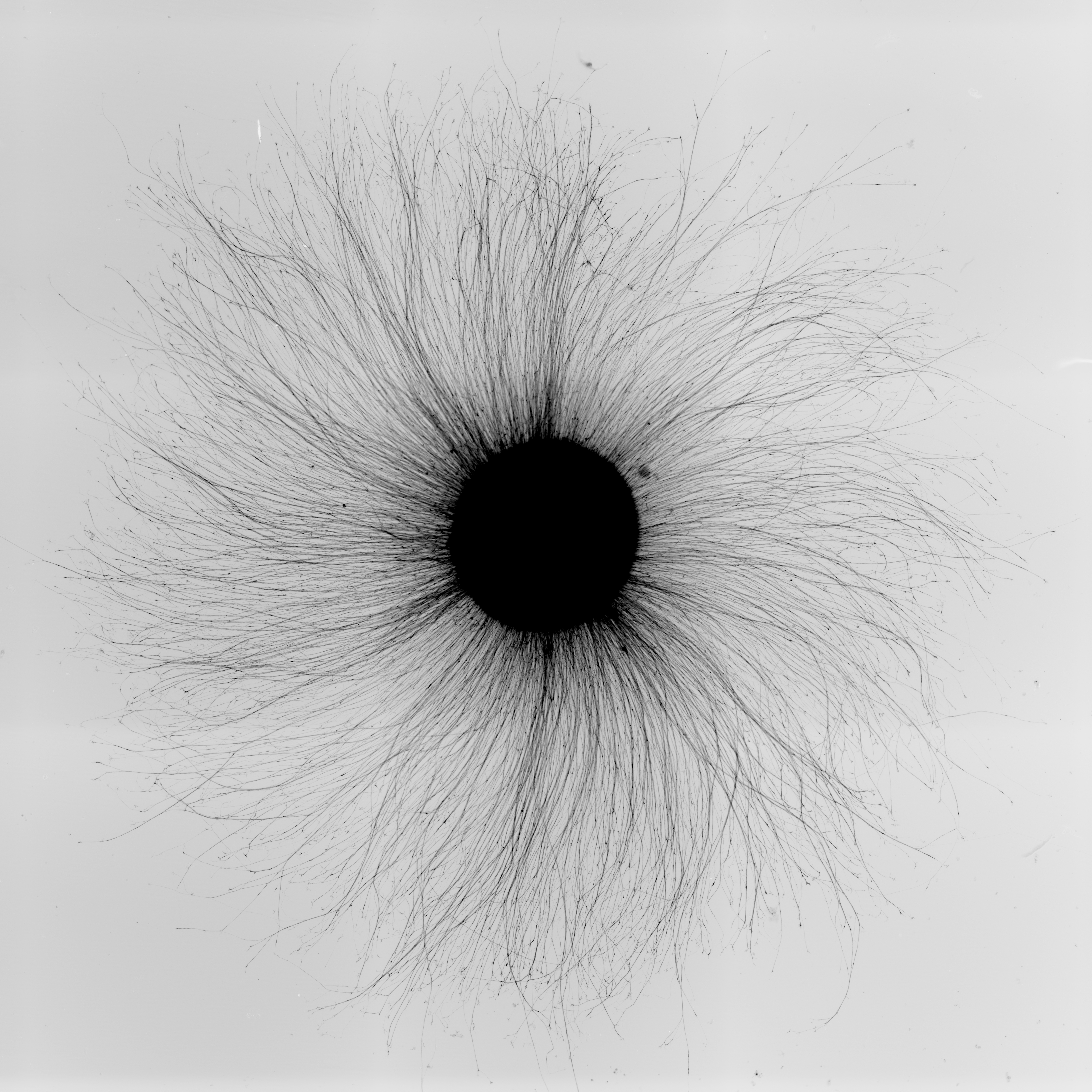
Pathways of Possibility– Julien Cicero
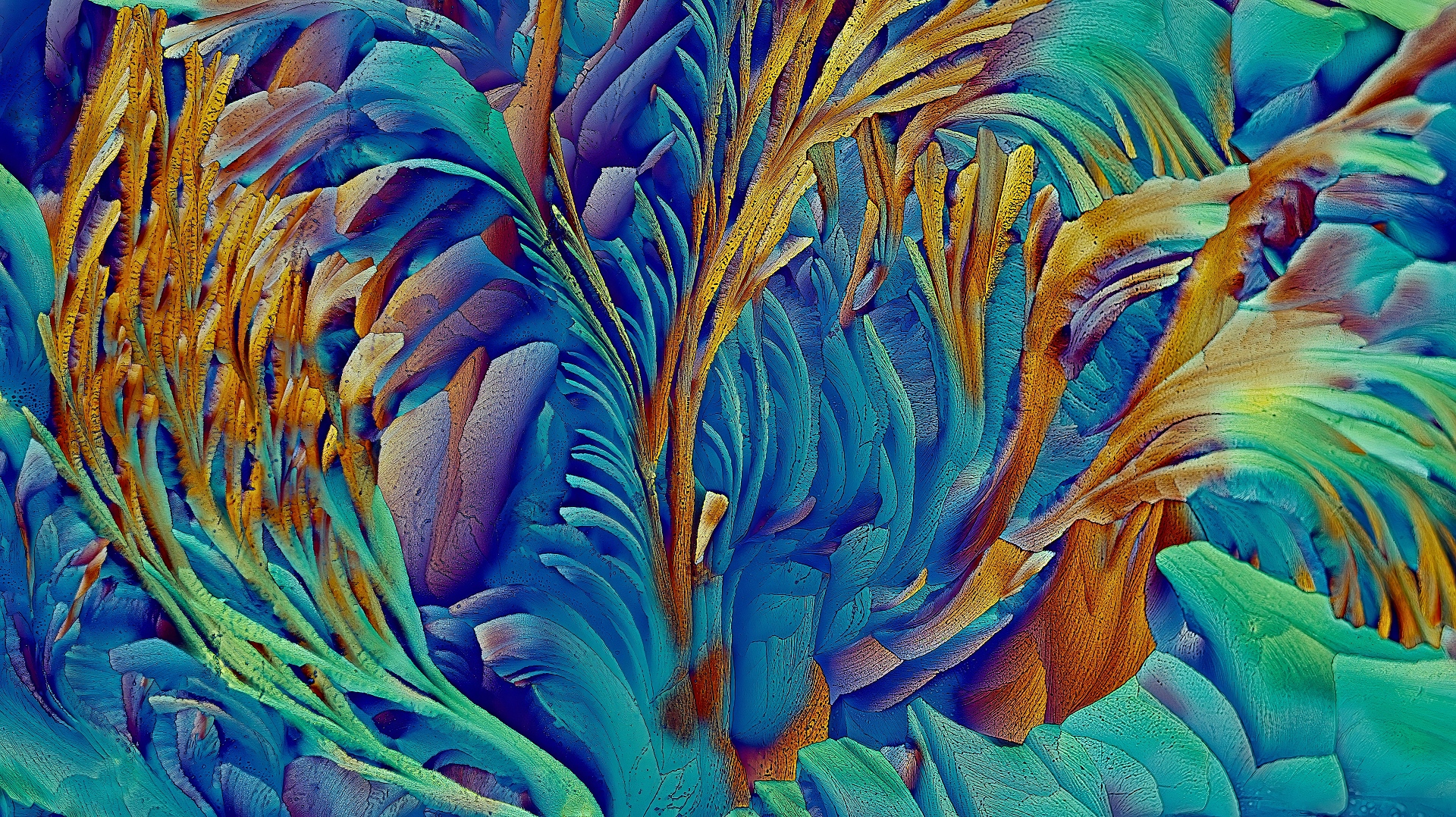

Crystallised B-alanine & l-Glutamine– Tian Olivier
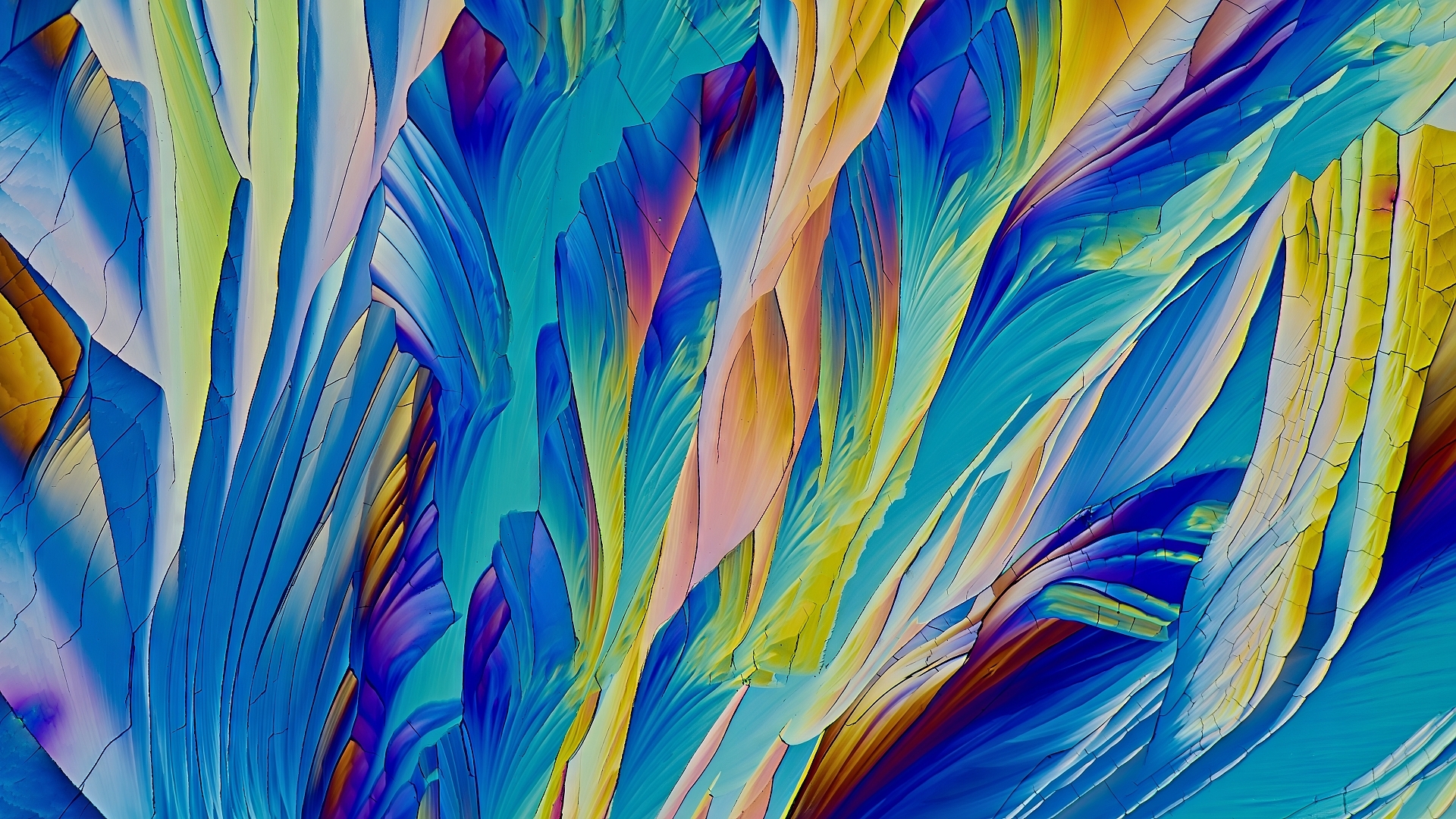

Crystallised B-Alanine and L-Glutamine– Tian Olivier
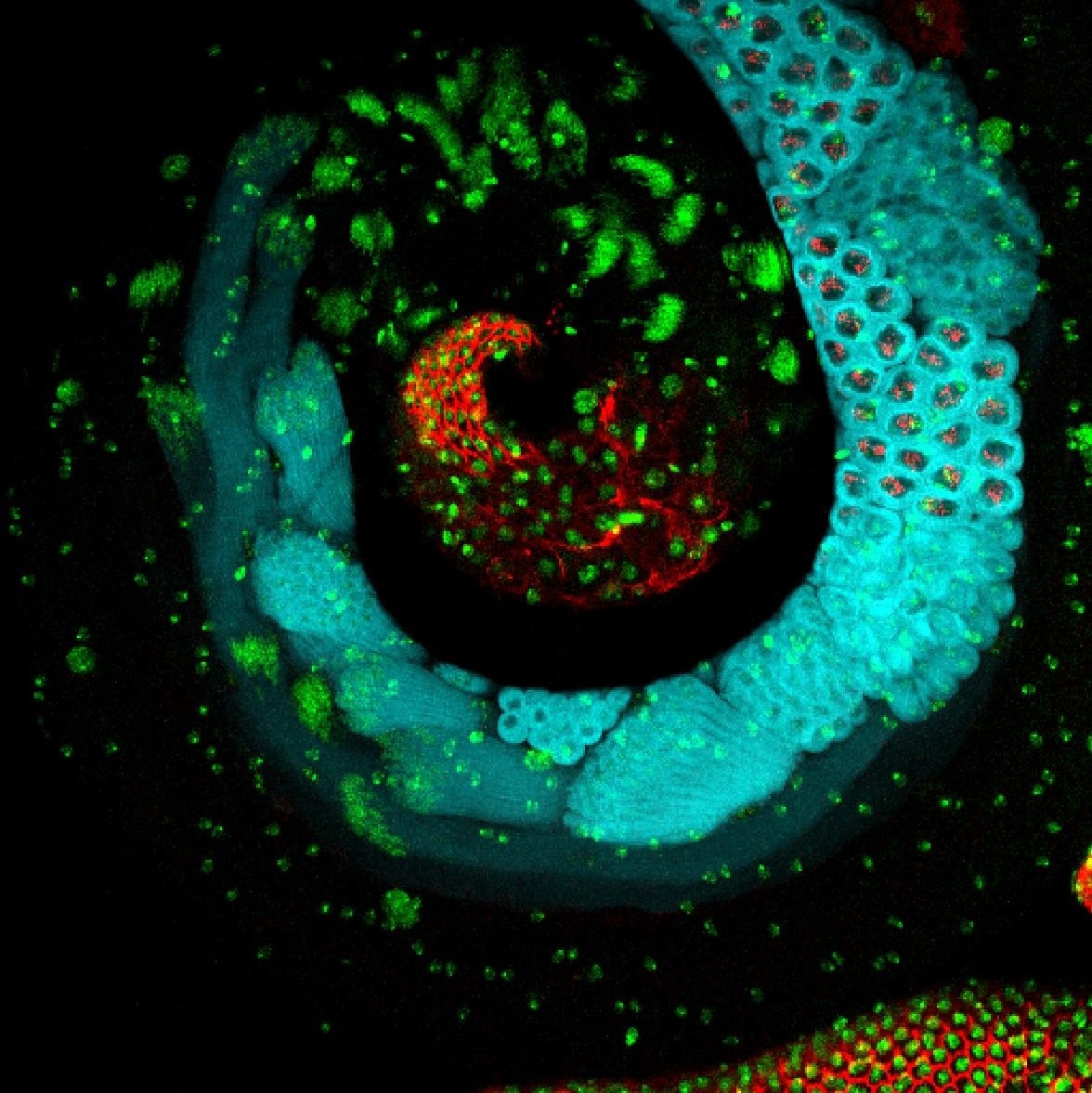
Festivus– Travis D. Carney**

Amino Acid crystals– Tian Olivier
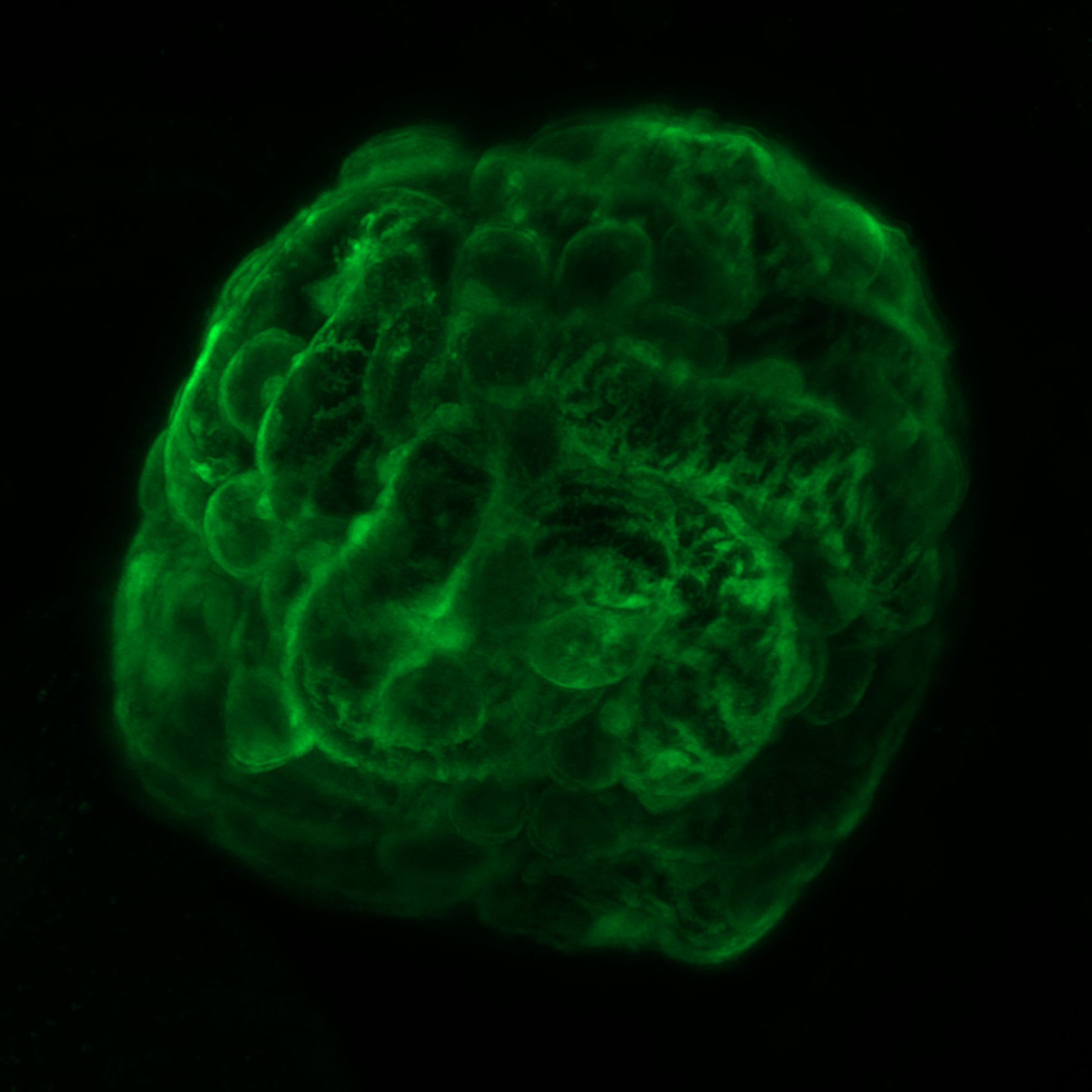
filtration ball– Caramai Kamei
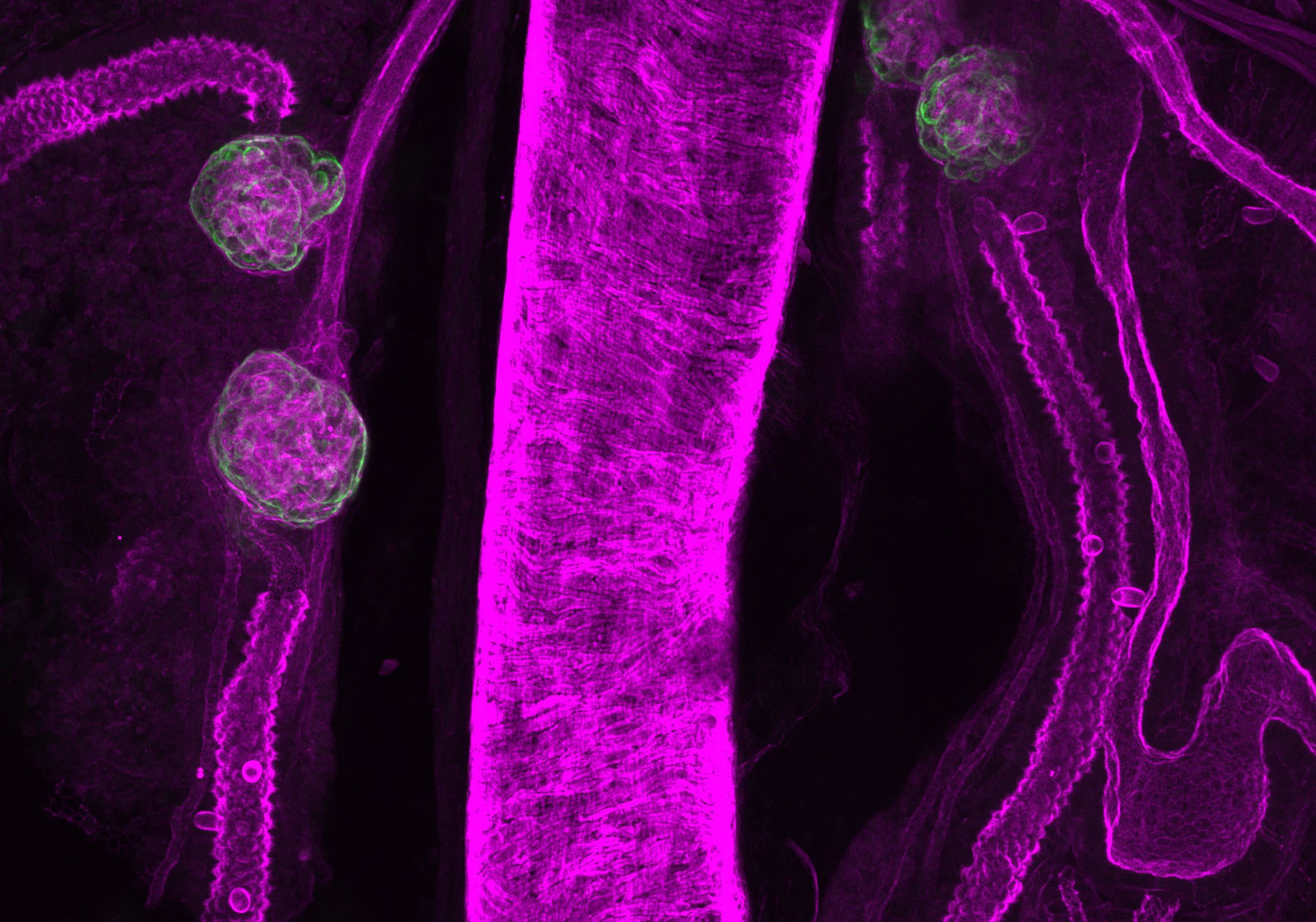
tree trunk- Caramai Kamei
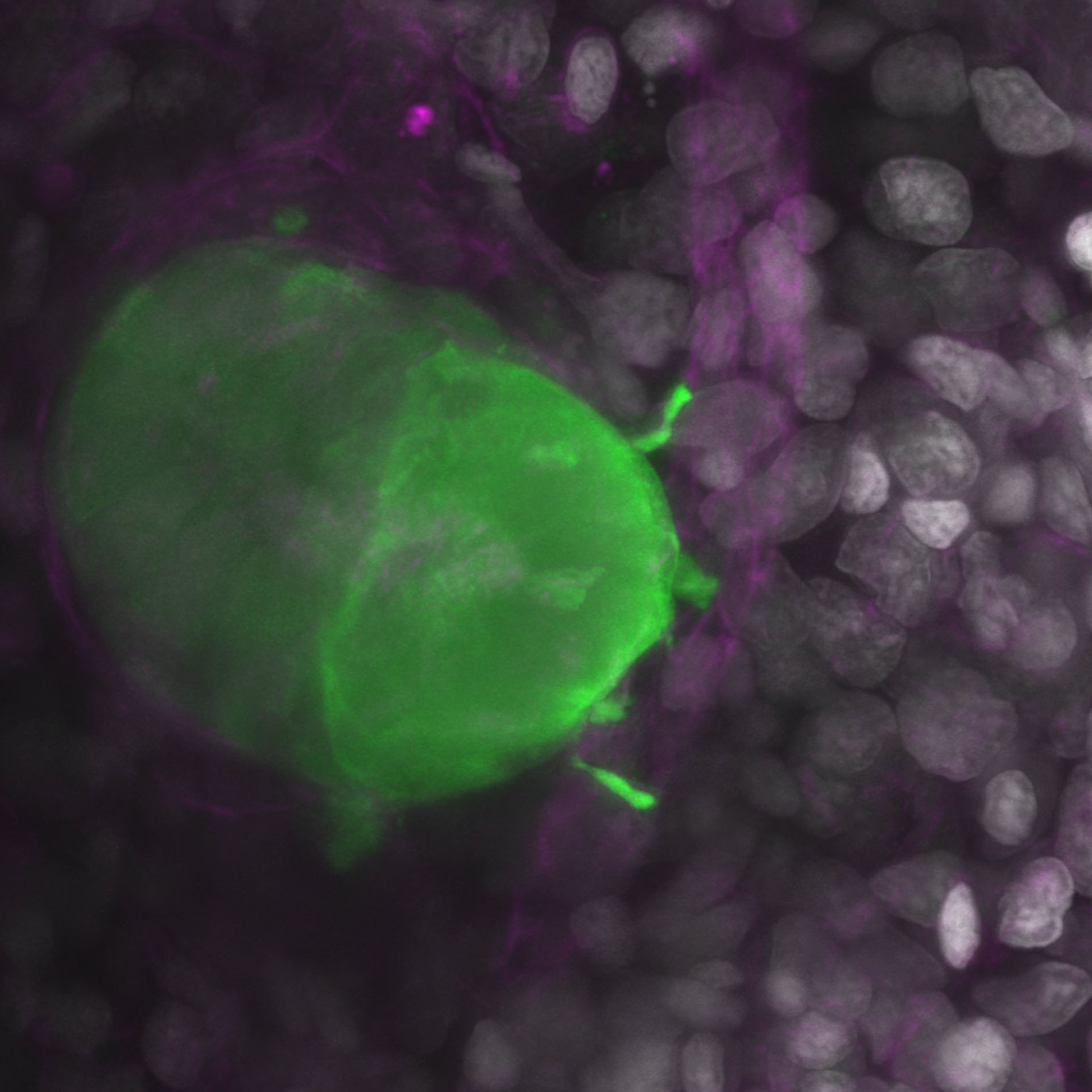
feelers– Caramai Kamei
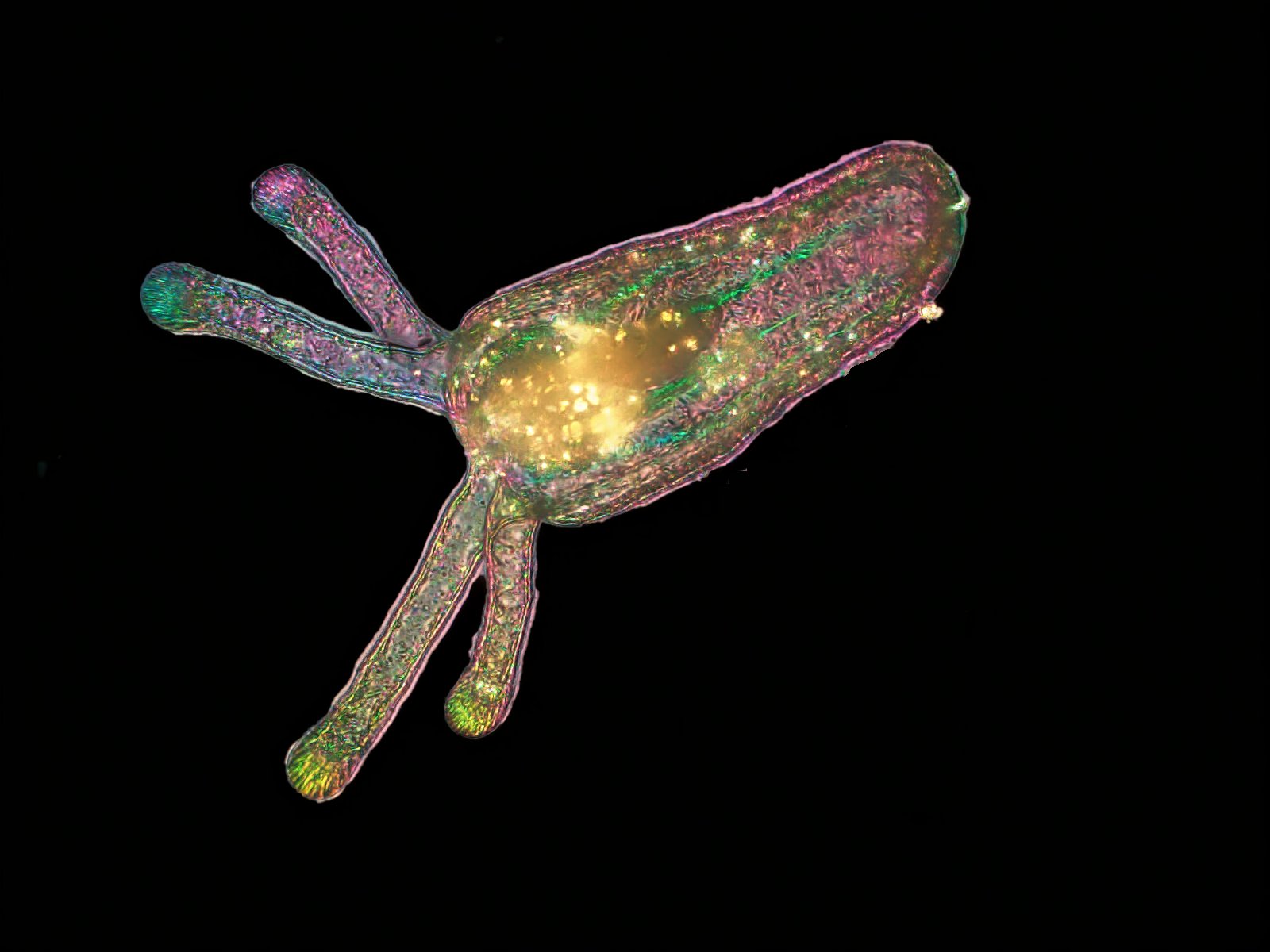
two-week-old larval sea anemone, Nematostella vectensis– Michael Shribak and Karen Echeverri
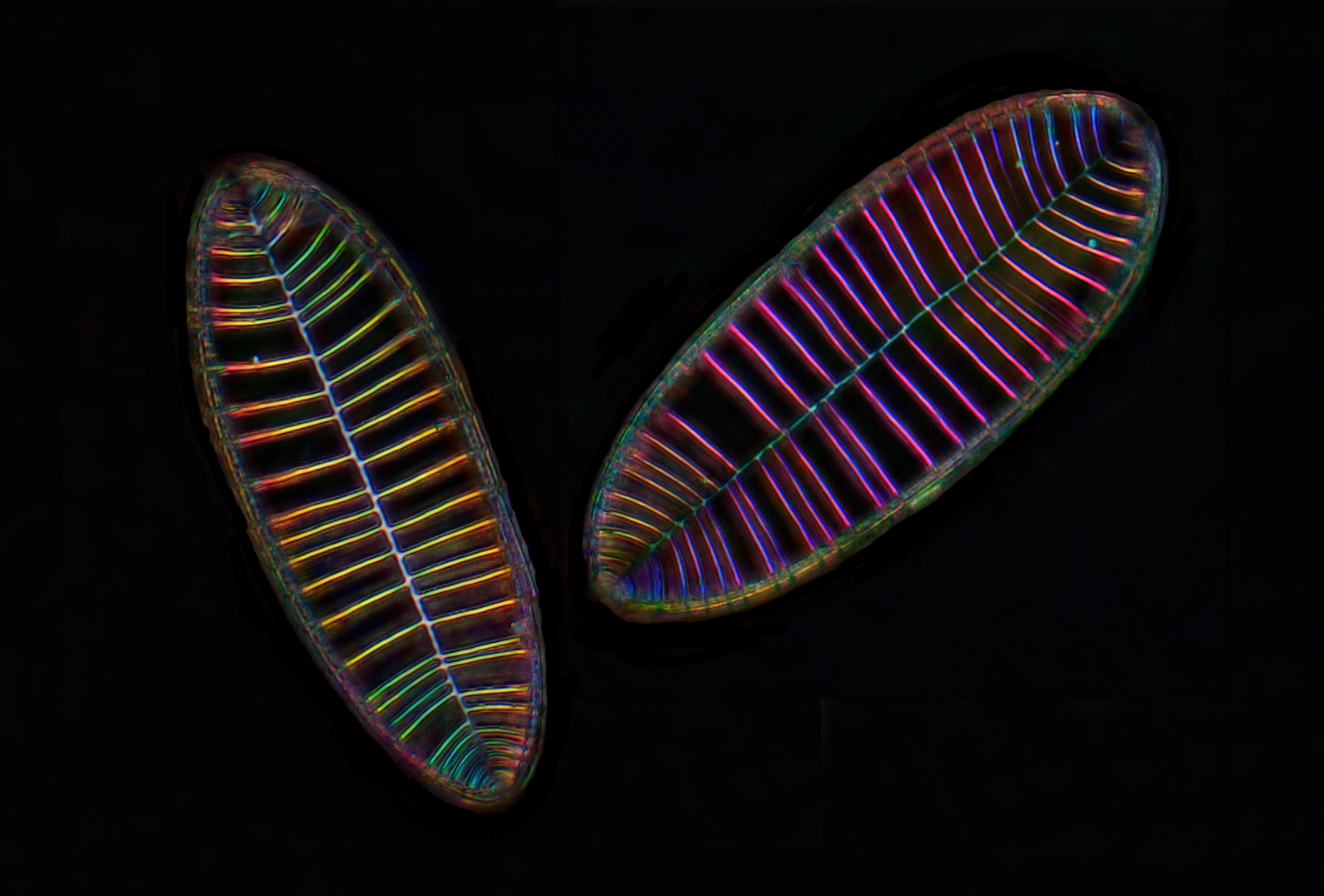
Diatom Surirella gemma- Michael Shribak

spider (Eriophora transmarina) silk – Michael Shribak

Neurons replicated…the University of Montreal logo in vitro!– Aurelie Stil
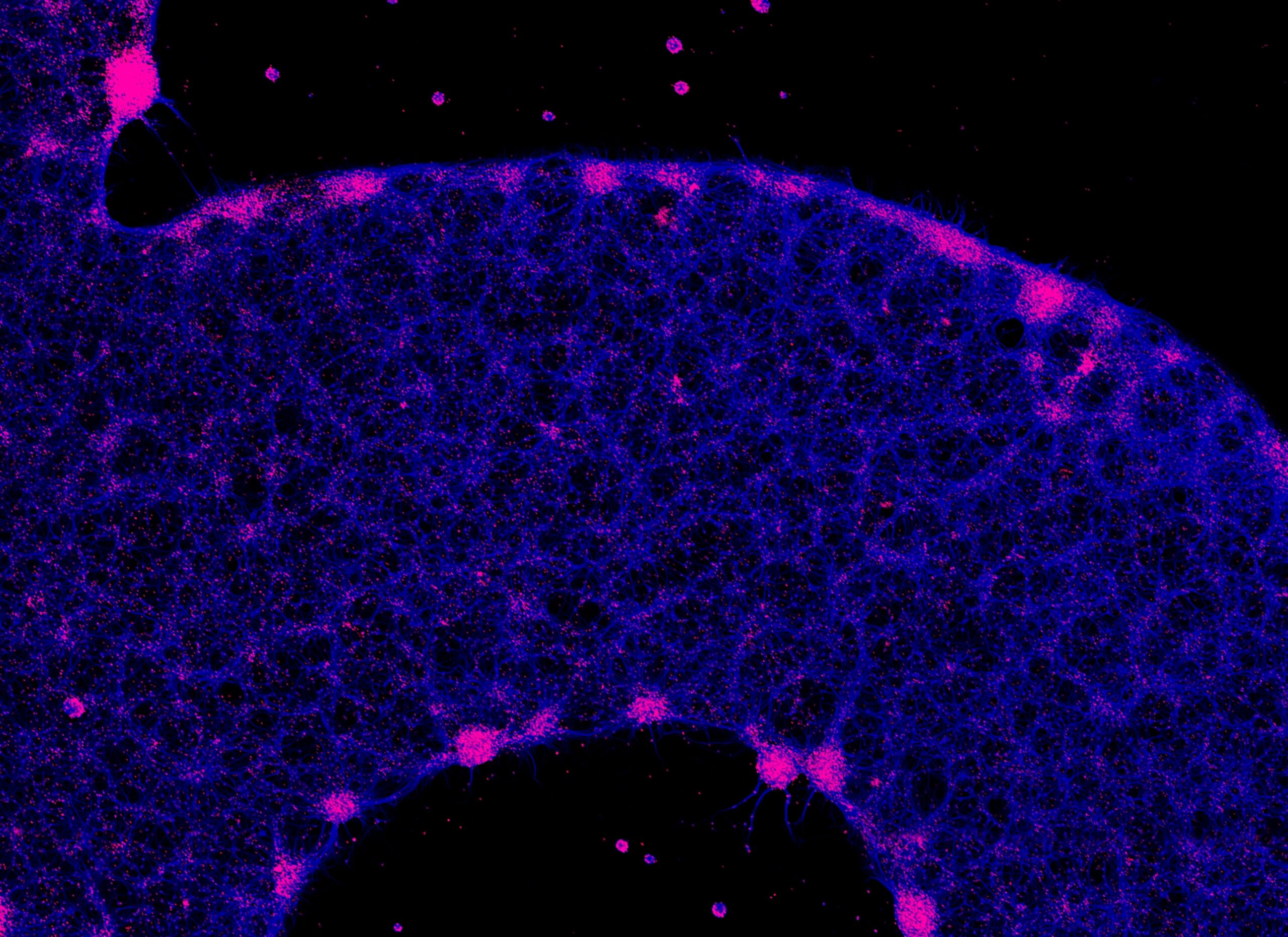
Neurons replicated…the University of Montreal logo in vitro!– Aurelie Stil
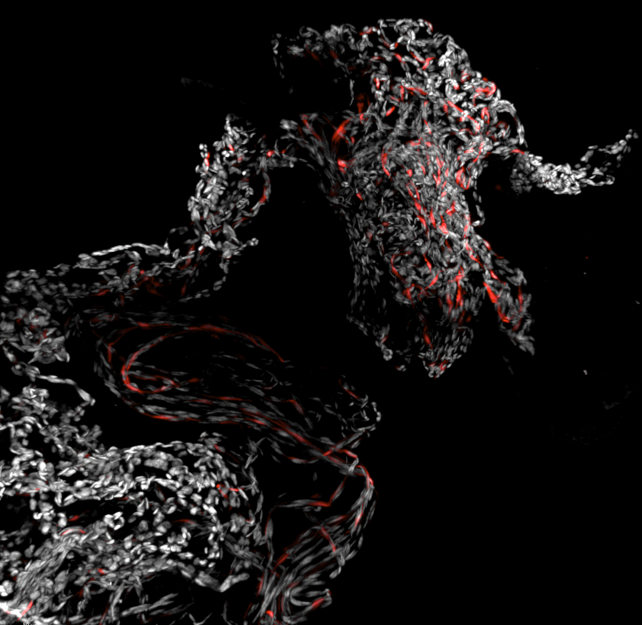
Sighting a racehorse– Team: Mariana De Niz, Jocelyn Salvador, Luisa Iruela-Arispe
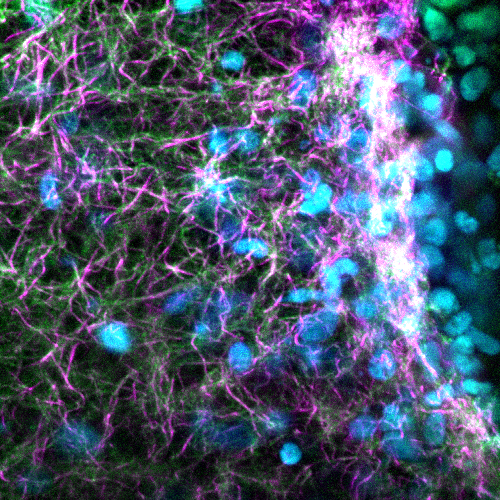
Fibrous Frontier– Jordan Miner
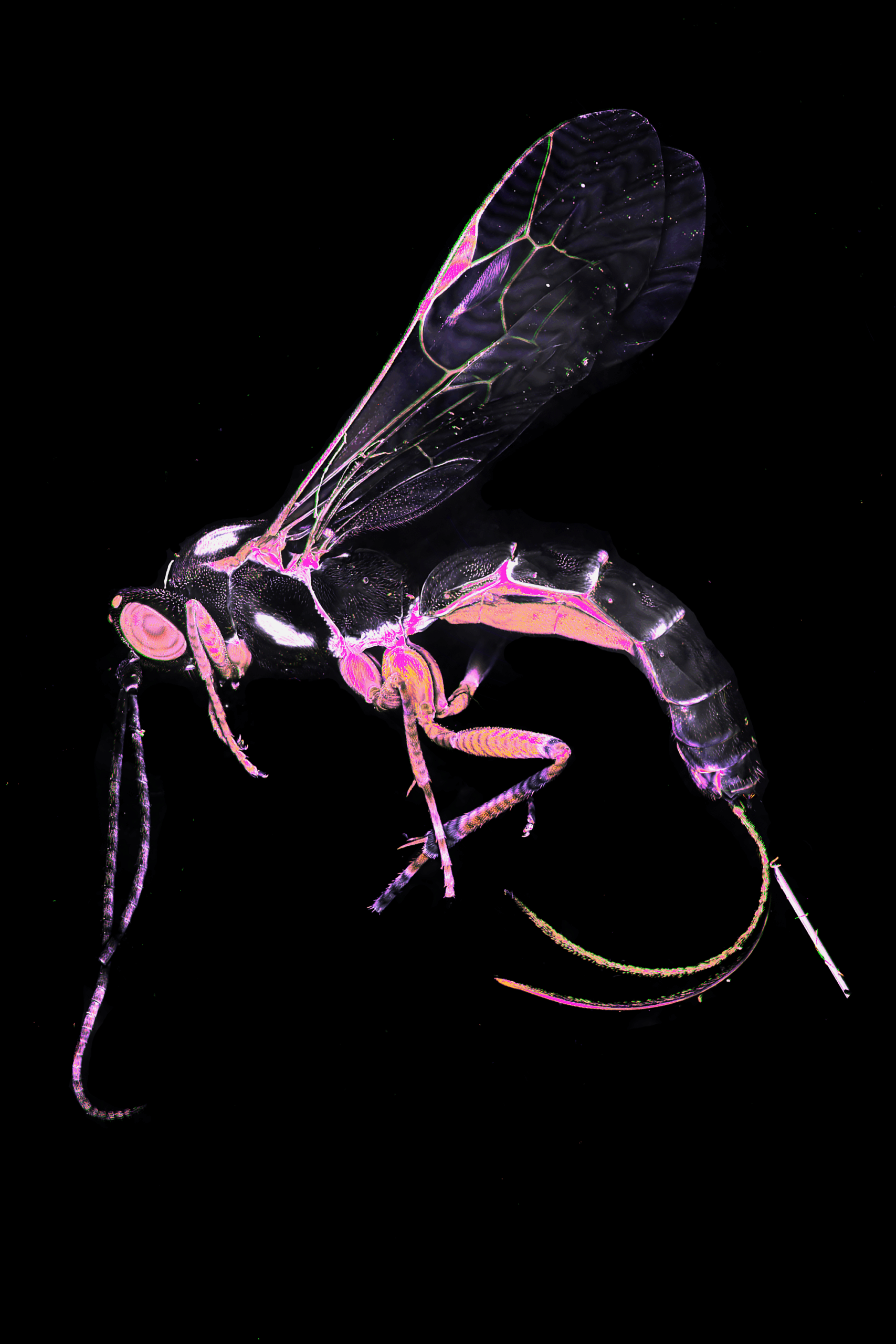

Fluorescent Flight: Parasitic Wasp in Pink Spectrum– Frederic Bonnet
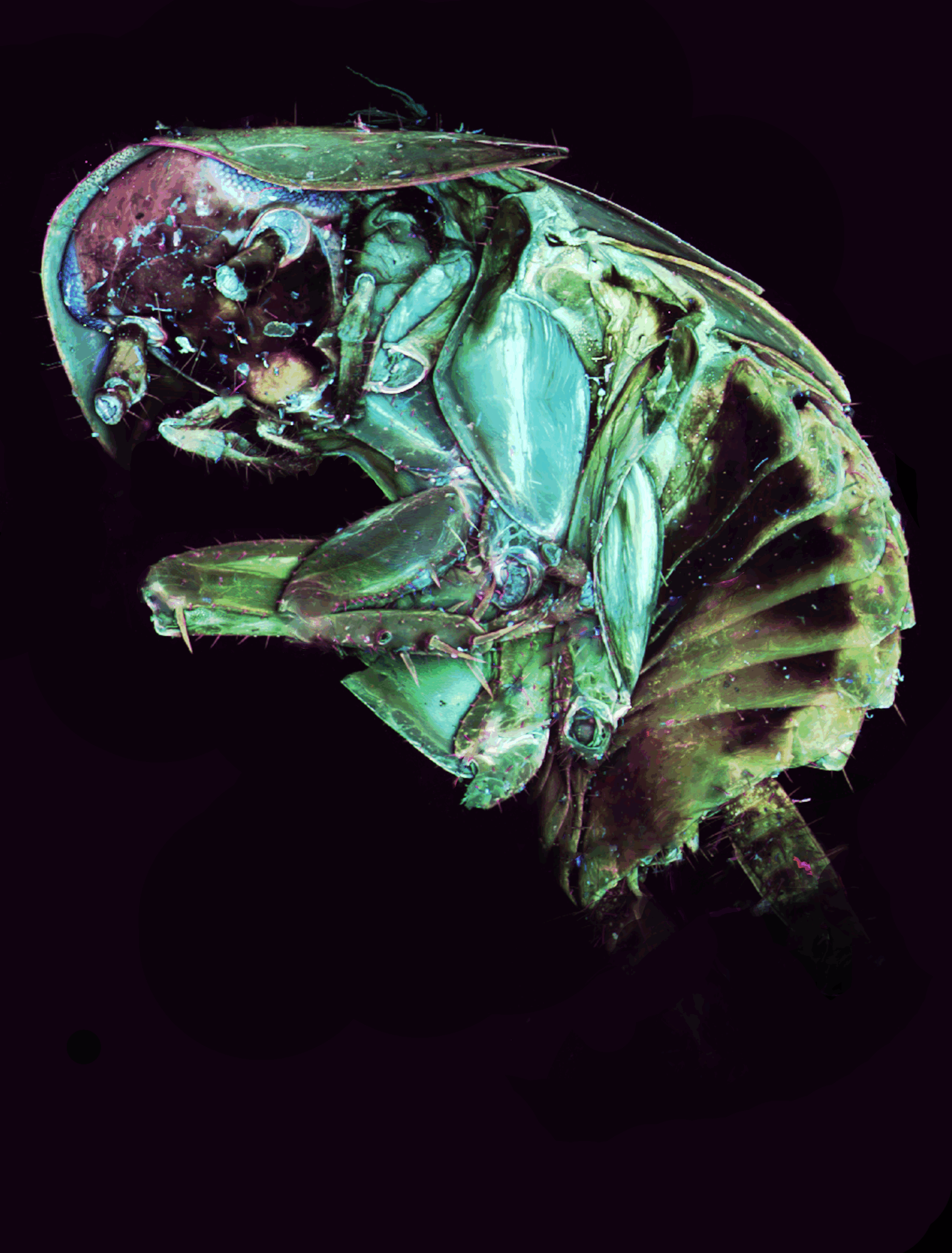

Emerald Architecture: Beetle’s Ventral Symphony– Frederic Bonnet
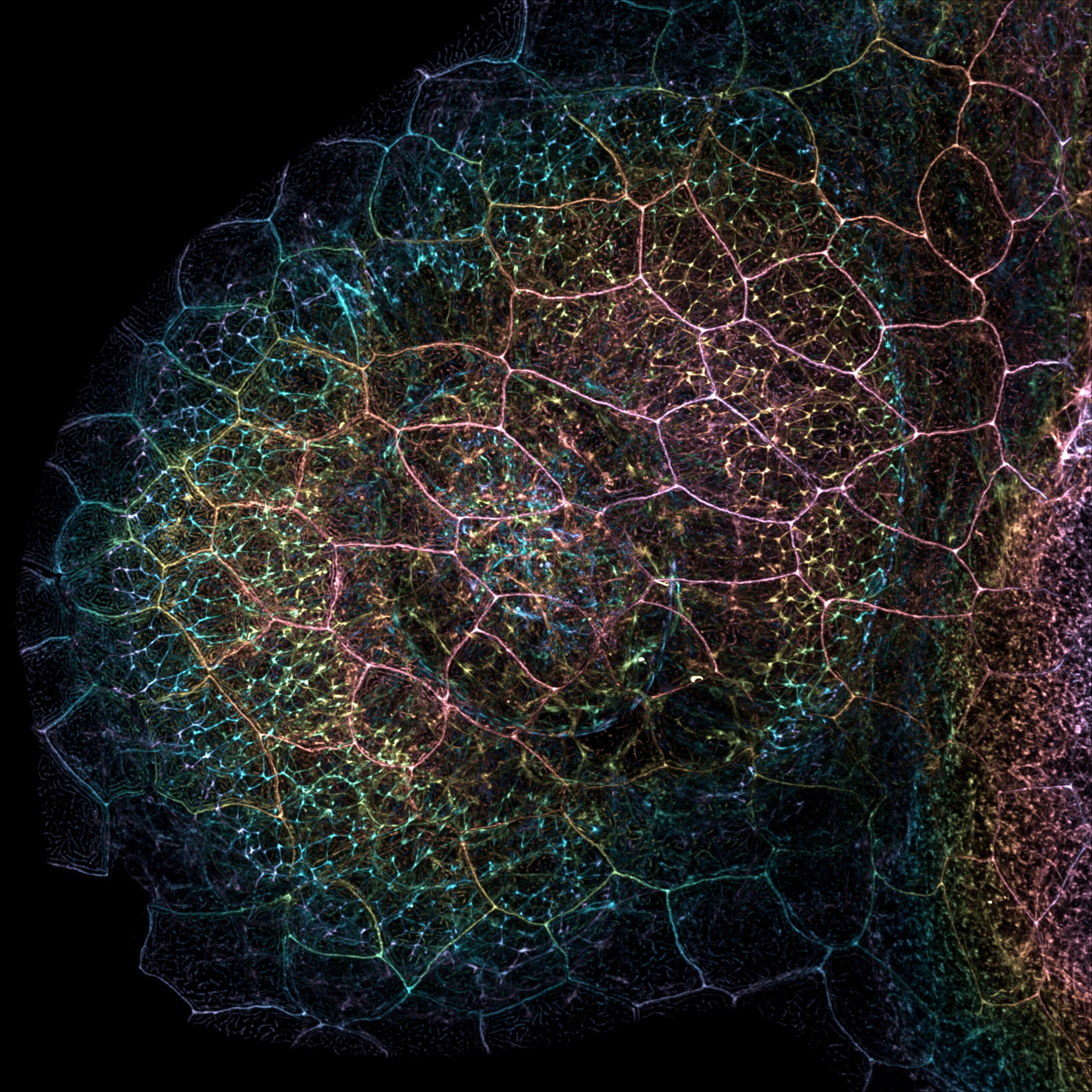
More than meets the eye: Actin in the zebrafish eye region- Caroline Hoppe, Valerie Tornini
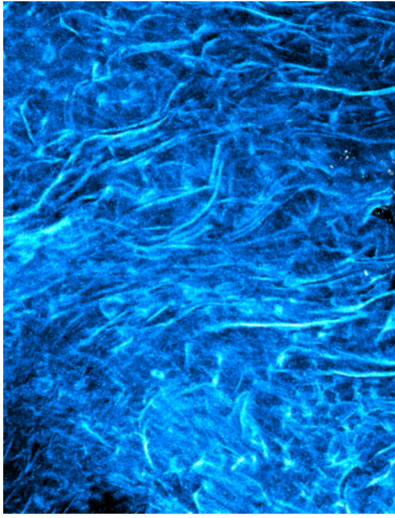
Harmonic Waves– Team: Mariana De Niz, Dina Arvanitis, Jocelyn Salvador, Luisa Iruela-Arispe
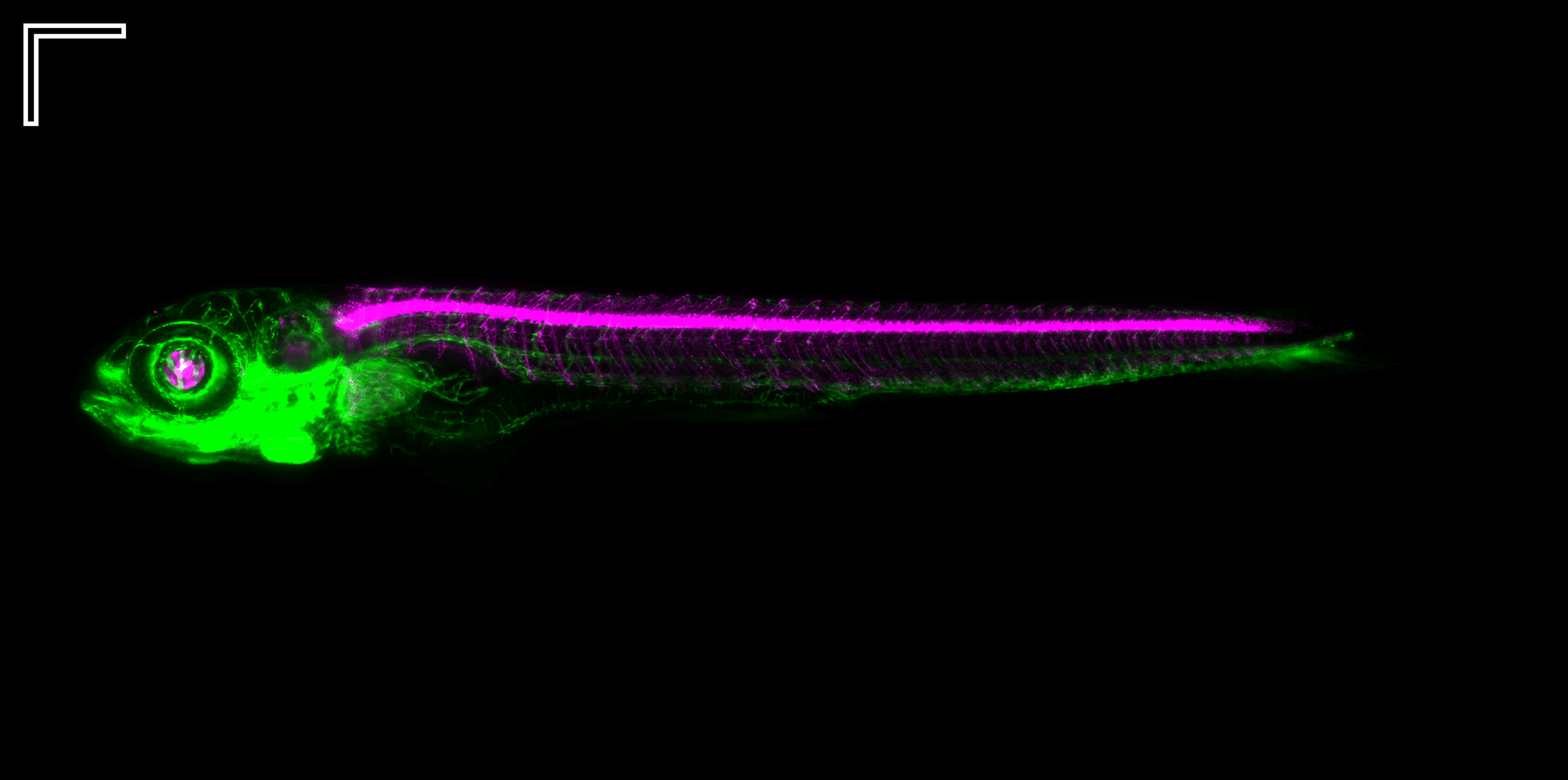
The zebrafish– Romain Menard
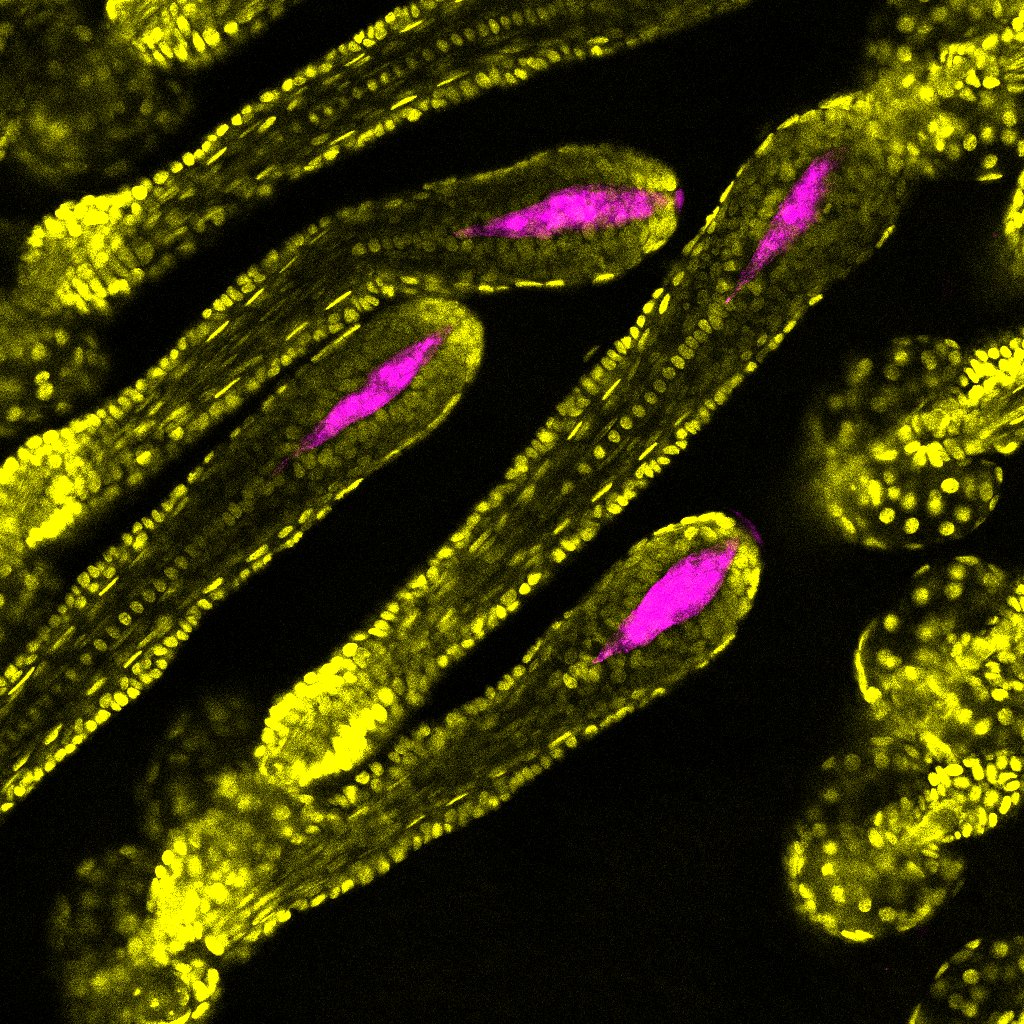
Murine Hair Follicles– Adrian Kwiatkowski
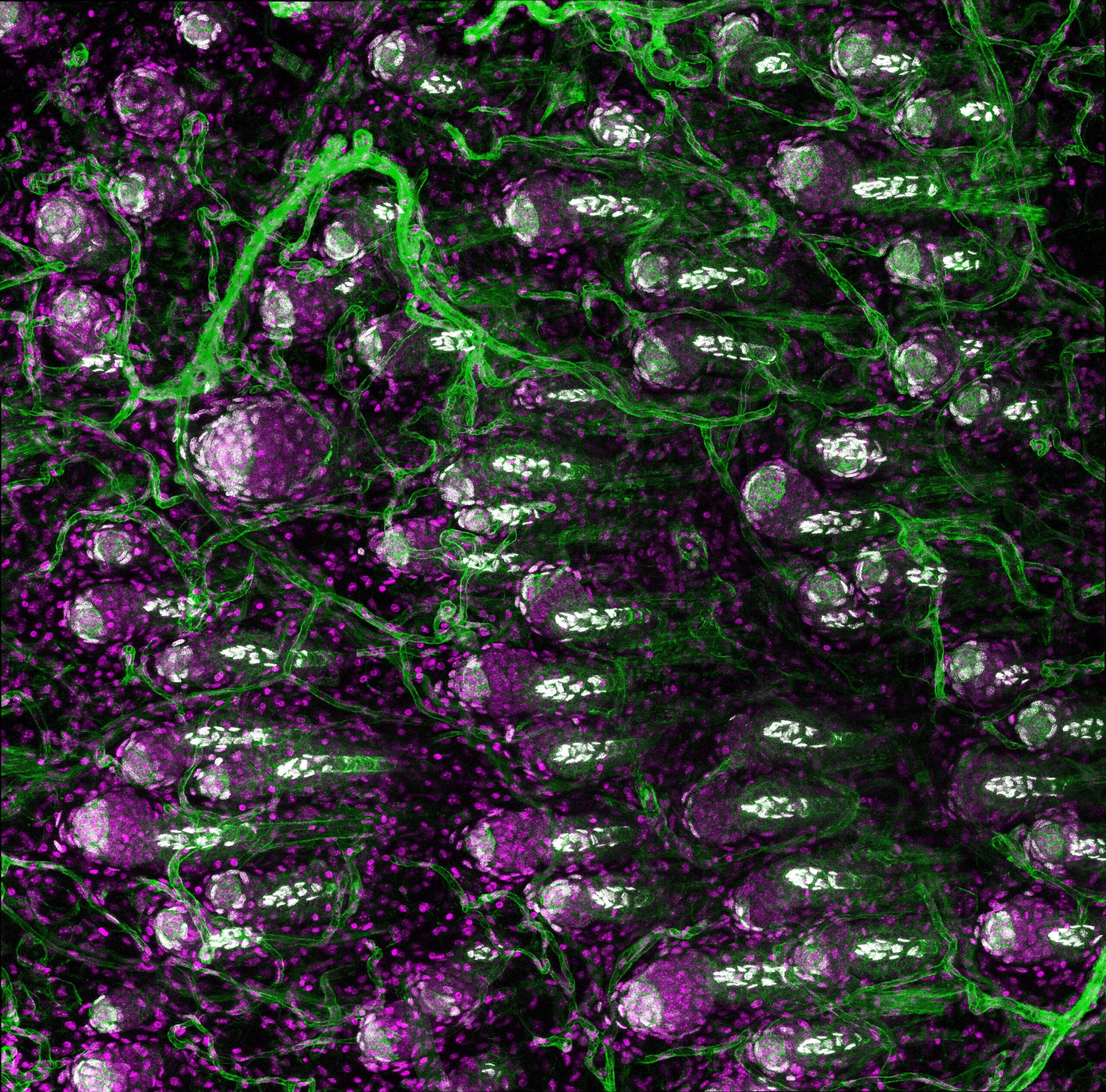
The hidden neighborhood in skin–Adrian Kwiatkowski
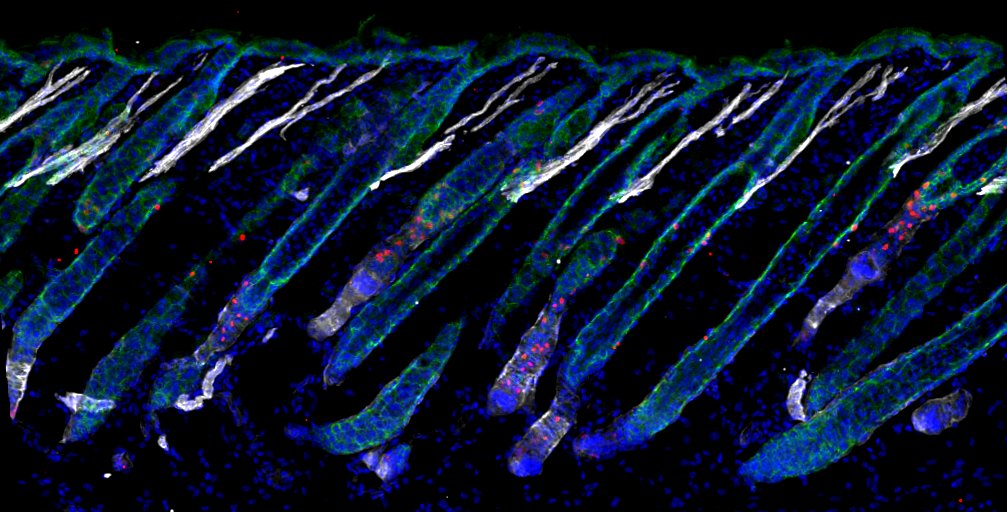
Regressing Hair Follicles in Murine Skin– Adrian Kwiatkowski
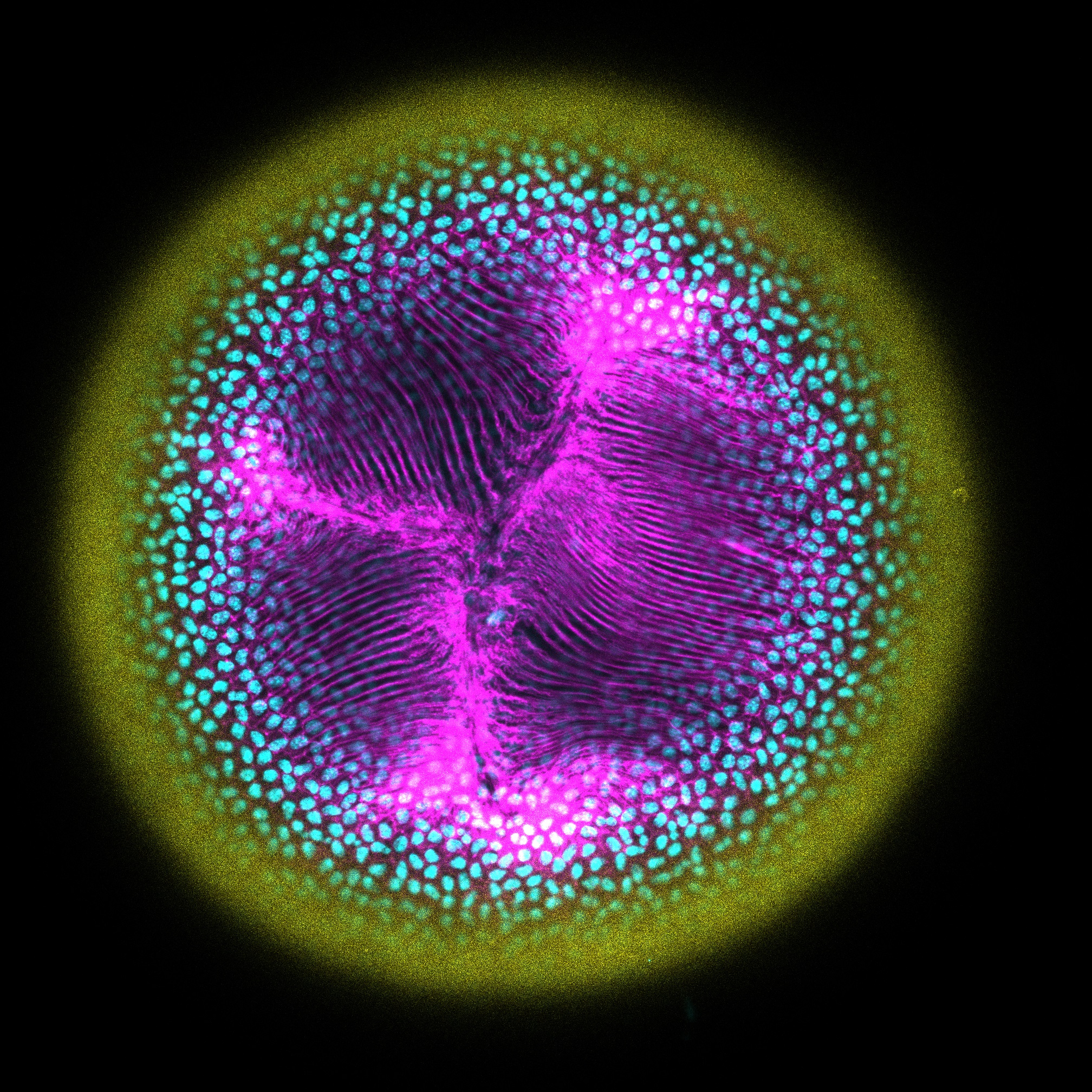
Where Cells Meet: The Lens Anterior Suture– Sepideh Cheheltani